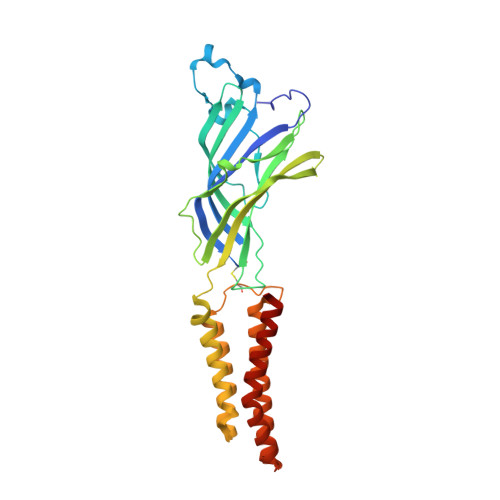
Structural Basis for Xenon Inhibition in a Cationic Pentameric Ligand-Gated Ion Channel.
Sauguet, L., Fourati, Z., Prange, T., Delarue, M., Colloc'h, N.(2016) PLoS One 11: e0149795-e0149795
- PubMed: 26910105 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0149795
- Primary Citation Related Structures:
4ZZB, 4ZZC - PubMed Abstract:
GLIC receptor is a bacterial pentameric ligand-gated ion channel whose action is inhibited by xenon. Xenon has been used in clinical practice as a potent gaseous anaesthetic for decades, but the molecular mechanism of interactions with its integral membrane receptor targets remains poorly understood. Here we characterize by X-ray crystallography the xenon-binding sites within both the open and "locally-closed" (inactive) conformations of GLIC. Major binding sites of xenon, which differ between the two conformations, were identified in three distinct regions that all belong to the trans-membrane domain of GLIC: 1) in an intra-subunit cavity, 2) at the interface between adjacent subunits, and 3) in the pore. The pore site is unique to the locally-closed form where the binding of xenon effectively seals the channel. A putative mechanism of the inhibition of GLIC by xenon is proposed, which might be extended to other pentameric cationic ligand-gated ion channels.
- Unité de Dynamique Structurale des Macromolécules (UMR 3528 CNRS) Institut Pasteur, Paris, France.
Organizational Affiliation: