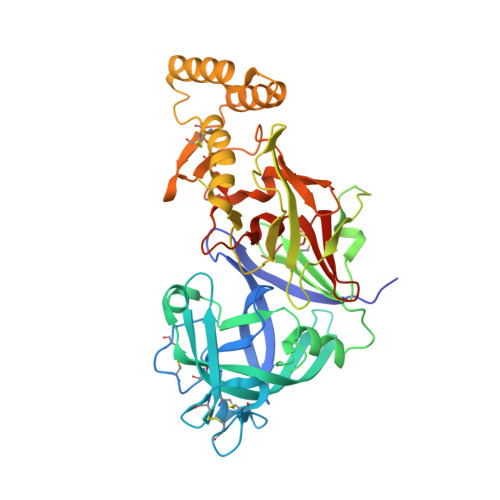
Structural basis for plasmepsin V inhibition that blocks export of malaria proteins to human erythrocytes.
Hodder, A.N., Sleebs, B.E., Czabotar, P.E., Gazdik, M., Xu, Y., O'Neill, M.T., Lopaticki, S., Nebl, T., Triglia, T., Smith, B.J., Lowes, K., Boddey, J.A., Cowman, A.F.(2015) Nat Struct Mol Biol 22: 590-596
- PubMed: 26214367 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3061
- Primary Citation Related Structures:
4ZL4 - PubMed Abstract:
Plasmepsin V, an essential aspartyl protease of malaria parasites, has a key role in the export of effector proteins to parasite-infected erythrocytes. Consequently, it is an important drug target for the two most virulent malaria parasites of humans, Plasmodium falciparum and Plasmodium vivax. We developed a potent inhibitor of plasmepsin V, called WEHI-842, which directly mimics the Plasmodium export element (PEXEL). WEHI-842 inhibits recombinant plasmepsin V with a half-maximal inhibitory concentration of 0.2 nM, efficiently blocks protein export and inhibits parasite growth. We obtained the structure of P. vivax plasmepsin V in complex with WEHI-842 to 2.4-Å resolution, which provides an explanation for the strict requirements for substrate and inhibitor binding. The structure characterizes both a plant-like fold and a malaria-specific helix-turn-helix motif that are likely to be important in cleavage of effector substrates for export.
- 1] Walter and Eliza Hall Institute of Medical Research, Parkville, Victoria, Australia. [2] Department of Medical Biology, University of Melbourne, Parkville, Victoria, Australia.
Organizational Affiliation: