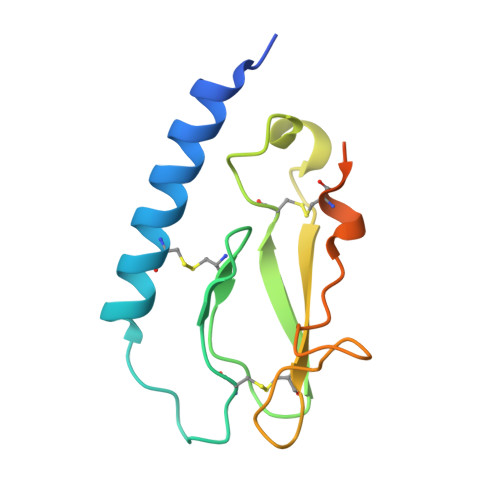
Discovery of the Once-Weekly Glucagon-Like Peptide-1 (GLP-1) Analogue Semaglutide.
Lau, J., Bloch, P., Schaffer, L., Pettersson, I., Spetzler, J., Kofoed, J., Madsen, K., Knudsen, L.B., McGuire, J., Steensgaard, D.B., Strauss, H.M., Gram, D.X., Knudsen, S.M., Nielsen, F.S., Thygesen, P., Reedtz-Runge, S., Kruse, T.(2015) J Med Chem 58: 7370-7380
- PubMed: 26308095 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00726
- Primary Citation Related Structures:
4ZGM - PubMed Abstract:
Liraglutide is an acylated glucagon-like peptide-1 (GLP-1) analogue that binds to serum albumin in vivo and is approved for once-daily treatment of diabetes as well as obesity. The aim of the present studies was to design a once weekly GLP-1 analogue by increasing albumin affinity and secure full stability against metabolic degradation. The fatty acid moiety and the linking chemistry to GLP-1 were the key features to secure high albumin affinity and GLP-1 receptor (GLP-1R) potency and in obtaining a prolonged exposure and action of the GLP-1 analogue. Semaglutide was selected as the optimal once weekly candidate. Semaglutide has two amino acid substitutions compared to human GLP-1 (Aib(8), Arg(34)) and is derivatized at lysine 26. The GLP-1R affinity of semaglutide (0.38 ± 0.06 nM) was three-fold decreased compared to liraglutide, whereas the albumin affinity was increased. The plasma half-life was 46.1 h in mini-pigs following i.v. administration, and semaglutide has an MRT of 63.6 h after s.c. dosing to mini-pigs. Semaglutide is currently in phase 3 clinical testing.
- Global Research, Novo Nordisk A/S , Novo Nordisk Park, DK-2760 Måløv, Denmark.
Organizational Affiliation: