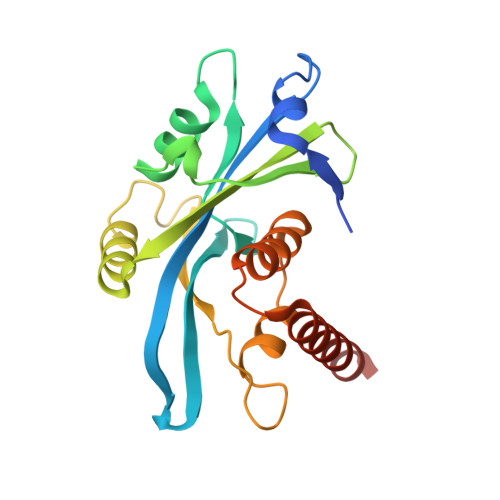
Crystal structure of syndesmos and its interaction with Syndecan-4 proteoglycan
Kim, H., Yoo, J., Lee, I., Kang, Y.J., Cho, H.S., Lee, W.(2015) Biochem Biophys Res Commun 463: 762-767
- PubMed: 26100207 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2015.06.010
- Primary Citation Related Structures:
4ZG0 - PubMed Abstract:
Syndesmos, nucleoside diphosphate linked moiety X (nudix)-type motif 16-like 1 (Nudt16l1), is evolutionarily divergent from the Nudt16 family. Syndesmos, which is co-localized with syndecan-4 cytoplasmic domain (Syn4(cyto)) in focal contacts, interacts with various cell adhesion adaptor proteins to control cell signaling. We determined the X-ray crystal structure of syndesmos; it is composed of seven α-helices and seven β-strands. Although syndesmos has a molecular topology similar to that of nudix hydrolase proteins, the structure of the nudix motif differs from that of X29. The dimeric interface of syndesmos is composed of α-helix 4, 7 and β-strand 2, 7, which primarily form hydrophobic interactions. The binding interaction between syndesmos and syn4(cyto) was characterized as a low-affinity interaction (Kd = 62 μM) by surface plasmon resonance (SPR) and nuclear magnetic resonance (NMR). The NMR resonances of Lys (177, 178, 179), Gly182, and Ser183 in the C1 region and Lys193 and Lys194 in the V region of syndecan-4 are perturbed upon syndesmos binding. Our results provide structural insight into the molecular function of syndesmos in the regulation of cell signaling via binding to syndecan-4.
- Department of Biochemistry, College of Life Science and Biotechnology, Yonsei University, Seoul, 120-749, South Korea.
Organizational Affiliation: