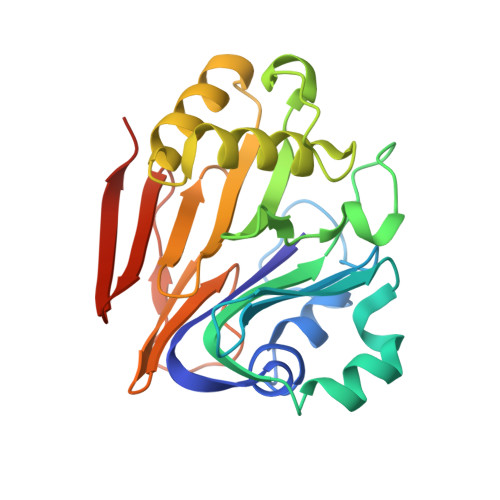
Structure of the Ergothioneine-Biosynthesis Amidohydrolase EgtC.
Vit, A., Mashabela, G.T., Blankenfeldt, W., Seebeck, F.P.(2015) Chembiochem 16: 1490-1496
- PubMed: 26079795 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201500168
- Primary Citation Related Structures:
4ZFJ, 4ZFK, 4ZFL - PubMed Abstract:
The ubiquitous sulfur metabolite ergothioneine is biosynthesized by oxidative attachment of a sulfur atom to the imidazole ring of Nα-trimethylhistidine. Most actinobacteria, including Mycobacterium tuberculosis, use γ-glutamyl cysteine as a sulfur donor. In subsequent steps the carbon scaffold of γ-glutamyl cysteine is removed by the glutamine amidohydrolase EgtC and the β-lyase EgtE. We determined the crystal structure of EgtC from Mycobacterium smegmatis in complex with its physiological substrate. The set of active site residues that define substrate specificity in EgtC are highly conserved, even in homologues that are not involved in ergothioneine production. This conservation is compounded by the phylogenetic distribution of EgtC-like enzymes indicates that their last common ancestor might have emerged for a purpose other than ergothioneine production.
- 3Structure and Function of Proteins, Helmholtz Centre for Infection Research, Inhoffenstrasse 7, 38124 Braunschweig (Germany).
Organizational Affiliation: