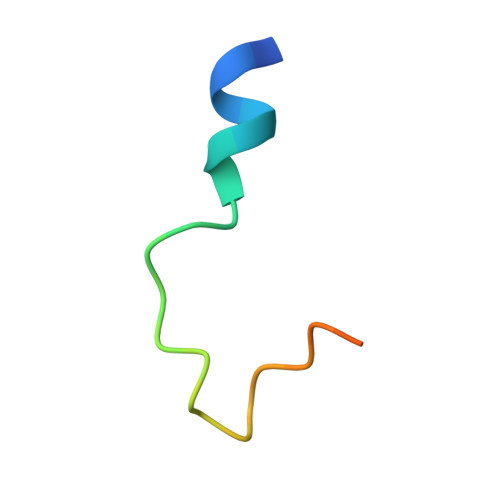
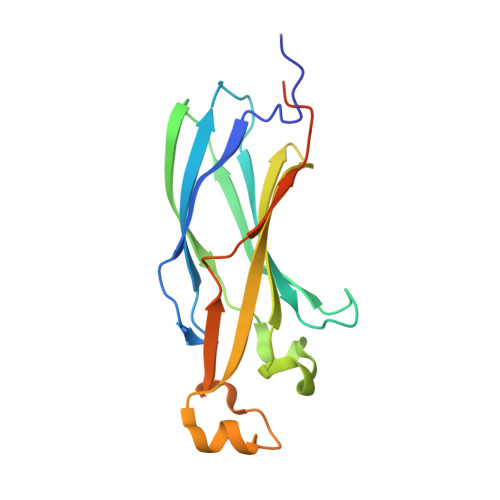
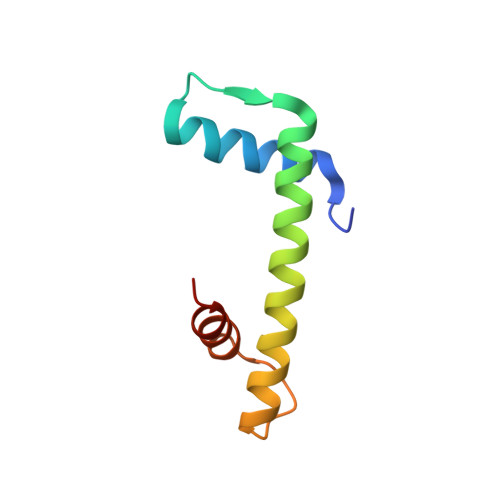
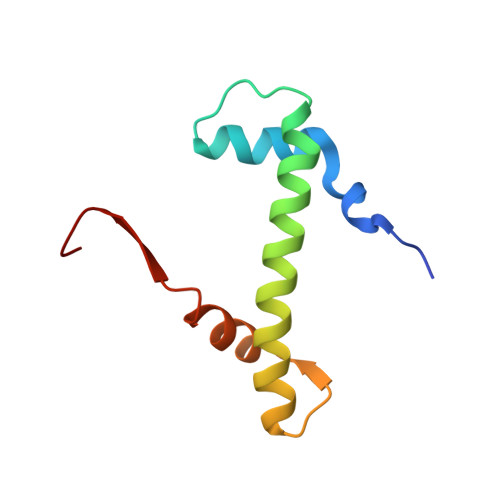
Ubinuclein-1 confers histone H3.3-specific-binding by the HIRA histone chaperone complex.
Daniel Ricketts, M., Frederick, B., Hoff, H., Tang, Y., Schultz, D.C., Singh Rai, T., Grazia Vizioli, M., Adams, P.D., Marmorstein, R.(2015) Nat Commun 6: 7711-7711
- PubMed: 26159857 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms8711
- Primary Citation Related Structures:
4ZBJ - PubMed Abstract:
Histone chaperones bind specific histones to mediate their storage, eviction or deposition from/or into chromatin. The HIRA histone chaperone complex, composed of HIRA, ubinuclein-1 (UBN1) and CABIN1, cooperates with the histone chaperone ASF1a to mediate H3.3-specific binding and chromatin deposition. Here we demonstrate that the conserved UBN1 Hpc2-related domain (HRD) is a novel H3.3-specific-binding domain. Biochemical and biophysical studies show the UBN1-HRD preferentially binds H3.3/H4 over H3.1/H4. X-ray crystallographic and mutational studies reveal that conserved residues within the UBN1-HRD and H3.3 G90 as key determinants of UBN1-H3.3-binding specificity. Comparison of the structure with the unrelated H3.3-specific chaperone DAXX reveals nearly identical points of contact between the chaperone and histone in the proximity of H3.3 G90, although the mechanism for H3.3 G90 recognition appears to be distinct. This study points to UBN1 as the determinant of H3.3-specific binding and deposition by the HIRA complex.
- 1] Department of Biochemistry and Biophysics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania 19104, USA [2] Graduate Group in Biochemistry and Molecular Biophysics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania 19104, USA.
Organizational Affiliation: