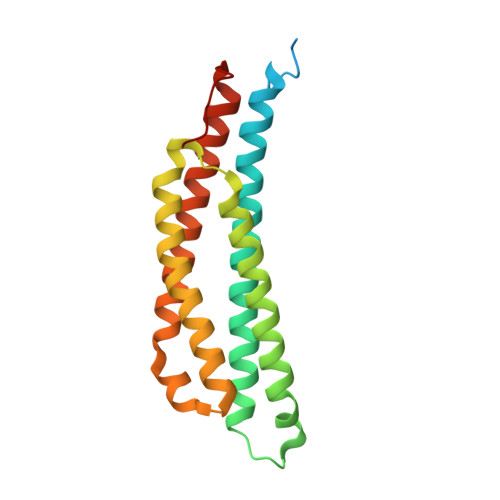
Structural Analysis of the Ligand-Binding Domain of the Aspartate Receptor Tar from Escherichia coli
Mise, T.(2016) Biochemistry 55: 3708-3713
- PubMed: 27292793 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.6b00160
- Primary Citation Related Structures:
4Z9H, 4Z9I, 4Z9J - PubMed Abstract:
The Escherichia coli cell-surface aspartate receptor Tar mediates bacterial chemotaxis toward an attractant, aspartate (Asp), and away from a repellent, Ni(2+). These signals are transmitted from the extracellular region of Tar to the cytoplasmic region via the transmembrane domain. The mechanism by which extracellular signals are transmitted into the cell through conformational changes in Tar is predicted to involve a piston displacement of one of the α4 helices of the homodimer. To understand the molecular mechanisms underlying the induction of Tar activity by an attractant, the three-dimensional structures of the E. coli Tar periplasmic domain with and without bound aspartate, Asp-Tar and apo-Tar, respectively, were determined. Of the two ligand-binding sites, only one site was occupied, and it clearly showed the electron density of an aspartate. The slight changes in conformation and the electrostatic surface potential around the aspartate-binding site were observed. In addition, the presence of an aspartate stabilized residues Phe-150' and Arg-73. A pistonlike displacement of helix α4b' was also induced by aspartate binding as predicted by the piston model. Taken together, these small changes might be related to the induction of Tar activity and might disturb binding of the second aspartate to the second binding site in E. coli.
- 2-19-3 Misato, Okinawa-shi, Okinawa 904-2153, Japan.
Organizational Affiliation: