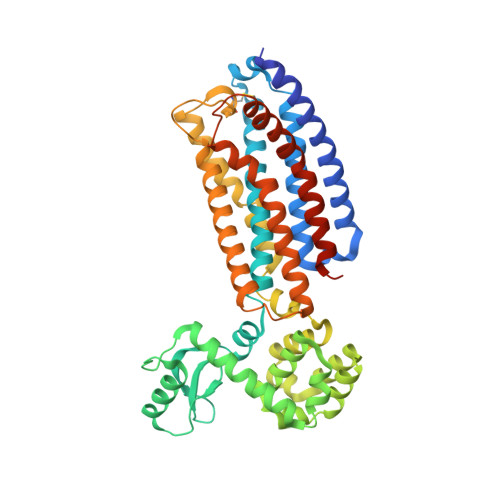
Decoding Corticotropin-Releasing Factor Receptor Type 1 Crystal Structures.
Dore, A.S., Bortolato, A., Hollenstein, K., Cheng, R.K.Y., Read, R.J., Marshall, F.H.(2017) Curr Mol Pharmacol 10: 334-344
- PubMed: 28183242 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.2174/1874467210666170110114727
- Primary Citation Related Structures:
4Z9G - PubMed Abstract:
The structural analysis of class B G protein-coupled receptors (GPCR), cell surface proteins responding to peptide hormones, has until recently been restricted to the extracellular domain (ECD). Corticotropin-releasing factor receptor type 1 (CRF1R) is a class B receptor mediating stress response and also considered a drug target for depression and anxiety. Here we report the crystal structure of the transmembrane domain of human CRF1R in complex with the small-molecule antagonist CP-376395 in a hexagonal setting with translational non-crystallographic symmetry. Molecular dynamics and metadynamics simulations on this novel structure and the existing TMD structure for CRF1R provides insight as to how the small molecule ligand gains access to the induced-fit allosteric binding site with implications for the observed selectivity against CRF2R. Furthermore, molecular dynamics simulations performed using a full-length receptor model point to key interactions between the ECD and extracellular loop 3 of the TMD providing insight into the full inactive state of multidomain class B GPCRs.
- Heptares Therapeutics Ltd, BioPark, Broadwater Road, Welwyn Garden City, Herts, AL7 3AX. United Kingdom.
Organizational Affiliation: