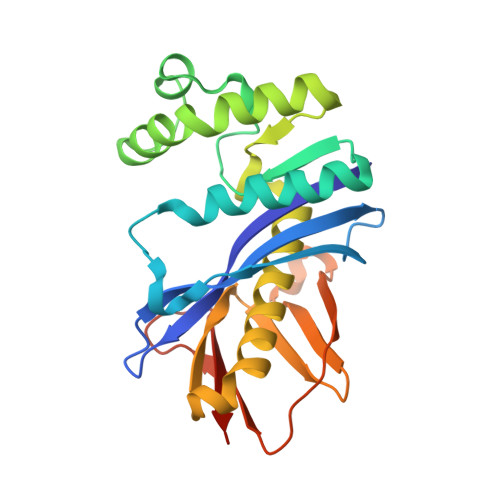
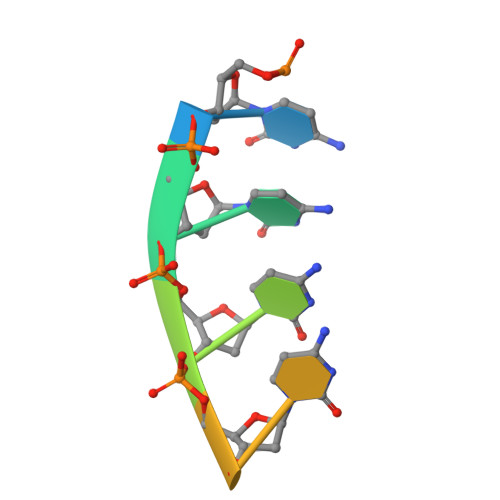
Structural basis for substrate recognition and processive cleavage mechanisms of the trimeric exonuclease PhoExo I
Miyazono, K., Ishino, S., Tsutsumi, K., Ito, T., Ishino, Y., Tanokura, M.(2015) Nucleic Acids Res 43: 7122-7136
- PubMed: 26138487 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkv654
- Primary Citation Related Structures:
4YOR, 4YOT, 4YOU, 4YOV, 4YOW, 4YOX, 4YOY - PubMed Abstract:
Nucleases play important roles in nucleic acid processes, such as replication, repair and recombination. Recently, we identified a novel single-strand specific 3'-5' exonuclease, PfuExo I, from the hyperthermophilic archaeon Pyrococcus furiosus, which may be involved in the Thermococcales-specific DNA repair system. PfuExo I forms a trimer and cleaves single-stranded DNA at every two nucleotides. Here, we report the structural basis for the cleavage mechanism of this novel exonuclease family. A structural analysis of PhoExo I, the homologous enzyme from P. horikoshii OT3, showed that PhoExo I utilizes an RNase H-like active site and possesses a 3'-OH recognition site ∼9 Å away from the active site, which enables cleavage at every two nucleotides. Analyses of the heterotrimeric and monomeric PhoExo I activities showed that trimerization is indispensable for its processive cleavage mechanism, but only one active site of the trimer is required.
- Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: