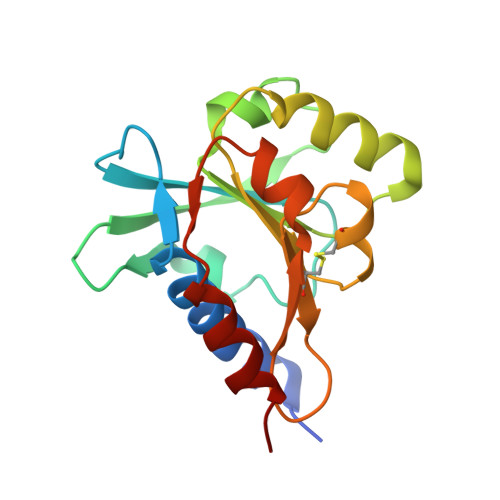
Crystal structures of YfiR from Pseudomonas aeruginosa in two redox states
Yang, X., Yang, X.-A., Xu, M., Zhou, L., Fan, Z., Jiang, T.(2015) Biochem Biophys Res Commun 461: 14-20
- PubMed: 25849887 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2015.03.160
- Primary Citation Related Structures:
4YN7, 4YN9, 4YNA - PubMed Abstract:
YfiBNR is a recently identified c-di-GMP regulatory system involved in bacterial biofilm formation. The periplasmic protein YfiR inhibits the diguanylate cyclase activity of the inner membrane protein YfiN, whereas YfiB in the outer membrane can release this inhibition by sequestration of YfiR. In addition, this system may respond to anoxic conditions via YfiR, although the detailed mechanism is still unknown. Here we report crystal structures of Pseudomonas aeruginosa YfiR in the absence and presence of oxidative glutathione. Our structures reveal the overall folding of YfiR for the first time and demonstrate that YfiR exist as a dimer. Comparison of the two structures in different redox states revealed a broken/formation of one disulfide bond (Cys71-Cys110) and local conformational change around the other one (Cys145-Cys152). Mutagenesis studies indicated that Cys145-Cys152 plays an important role in maintaining the correct folding of YfiR.
- Chinese Academy of Sciences Key Laboratory of Infection and Immunity, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, PR China.
Organizational Affiliation: