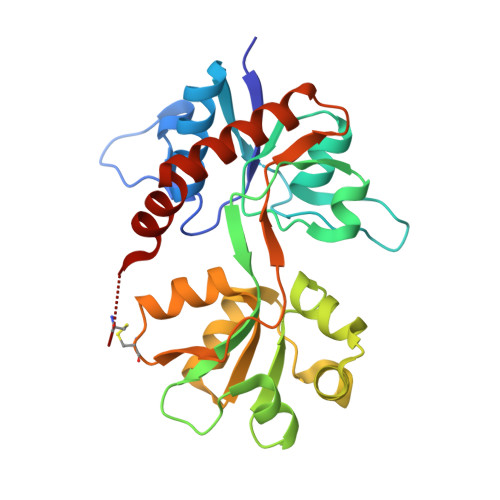
Structure-Activity Relationship Study of Ionotropic Glutamate Receptor Antagonist (2S,3R)-3-(3-Carboxyphenyl)pyrrolidine-2-carboxylic Acid.
Krogsgaard-Larsen, N., Storgaard, M., Moller, C., Demmer, C.S., Hansen, J., Han, L., Monrad, R.N., Nielsen, B., Tapken, D., Pickering, D.S., Kastrup, J.S., Frydenvang, K., Bunch, L.(2015) J Med Chem 58: 6131-6150
- PubMed: 26200741 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00750
- Primary Citation Related Structures:
4YMA, 4YMB - PubMed Abstract:
Herein we describe the first structure-activity relationship study of the broad-range iGluR antagonist (2S,3R)-3-(3-carboxyphenyl)pyrrolidine-2-carboxylic acid (1) by exploring the pharmacological effect of substituents in the 4, 4', or 5' positions and the bioisosteric substitution of the distal carboxylic acid for a phosphonic acid moiety. Of particular interest is a hydroxyl group in the 4' position 2a which induced a preference in binding affinity for homomeric GluK3 over GluK1 (Ki = 0.87 and 4.8 μM, respectively). Two X-ray structures of ligand binding domains were obtained: 2e in GluA2-LBD and 2f in GluK1-LBD, both at 1.9 Å resolution. Compound 2e induces a D1-D2 domain opening in GluA2-LBD of 17.3-18.8° and 2f a domain opening in GluK1-LBD of 17.0-17.5° relative to the structures with glutamate. The pyrrolidine-2-carboxylate moiety of 2e and 2f shows a similar binding mode as kainate. The 3-carboxyphenyl ring of 2e and 2f forms contacts comparable to those of the distal carboxylate in kainate.
- †Chemical Neuroscience Group, ‡Biostructural Research Group, §Medicinal Chemistry Group, ∥Molecular, Cellular Pharmacology Group, Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen, Universitetsparken 2, 2100 Copenhagen Ø, Denmark.
Organizational Affiliation: