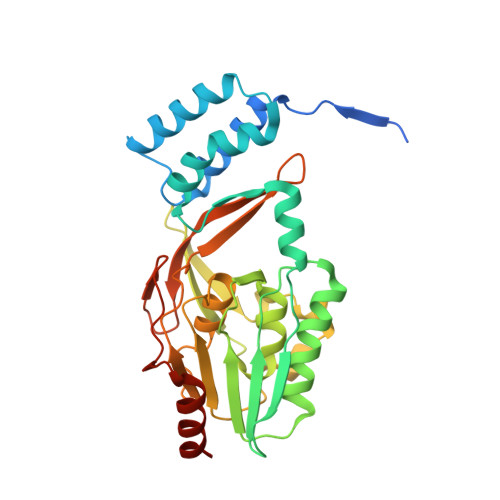
Structural basis of lantibiotic recognition by the nisin resistance protein from Streptococcus agalactiae.
Khosa, S., Frieg, B., Mulnaes, D., Kleinschrodt, D., Hoeppner, A., Gohlke, H., Smits, S.H.(2016) Sci Rep 6: 18679-18679
- PubMed: 26727488 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep18679
- Primary Citation Related Structures:
4Y68 - PubMed Abstract:
Lantibiotics are potent antimicrobial peptides. Nisin is the most prominent member and contains five crucial lanthionine rings. Some clinically relevant bacteria express membrane-associated resistance proteins that proteolytically inactivate nisin. However, substrate recognition and specificity of these proteins is unknown. Here, we report the first three-dimensional structure of a nisin resistance protein from Streptococcus agalactiae (SaNSR) at 2.2 Å resolution. It contains an N-terminal helical bundle, and protease cap and core domains. The latter harbors the highly conserved TASSAEM region, which lies in a hydrophobic tunnel formed by all domains. By integrative modeling, mutagenesis studies, and genetic engineering of nisin variants, a model of the SaNSR/nisin complex is generated, revealing that SaNSR recognizes the last C-terminally located lanthionine ring of nisin. This determines the substrate specificity of SaNSR and ensures the exact coordination of the nisin cleavage site at the TASSAEM region.
- Institute of Biochemistry, Heinrich Heine University, Universitätsstr. 1, 40225 Düsseldorf, Germany.
Organizational Affiliation: