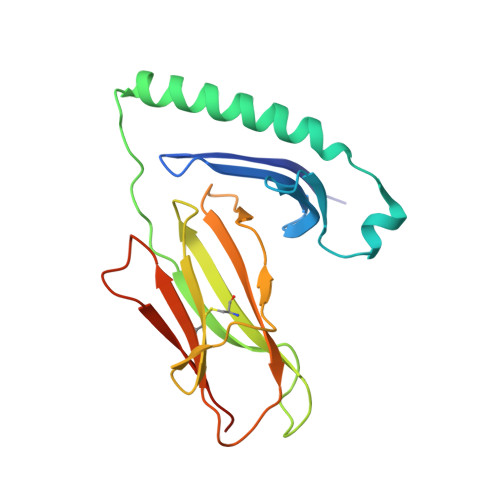
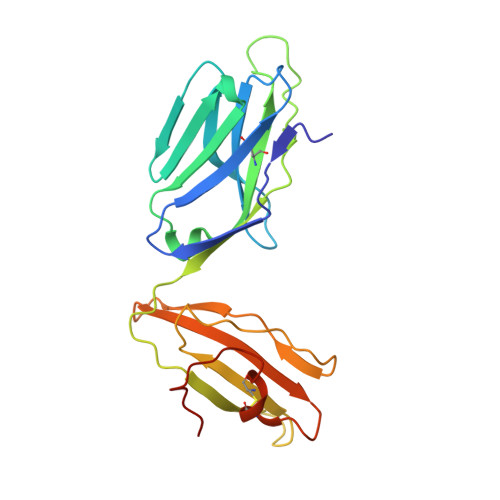
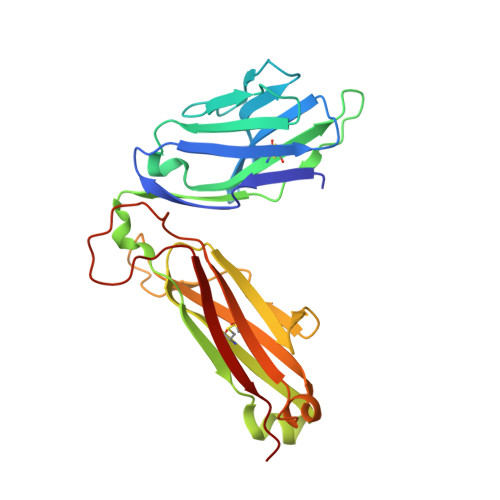
T cell receptor reversed polarity recognition of a self-antigen major histocompatibility complex.
Beringer, D.X., Kleijwegt, F.S., Wiede, F., van der Slik, A.R., Loh, K.L., Petersen, J., Dudek, N.L., Duinkerken, G., Laban, S., Joosten, A., Vivian, J.P., Chen, Z., Uldrich, A.P., Godfrey, D.I., McCluskey, J., Price, D.A., Radford, K.J., Purcell, A.W., Nikolic, T., Reid, H.H., Tiganis, T., Roep, B.O., Rossjohn, J.(2015) Nat Immunol 16: 1153-1161
- PubMed: 26437244 Search on PubMed
- DOI: https://doi.org/10.1038/ni.3271
- Primary Citation Related Structures:
4Y19, 4Y1A - PubMed Abstract:
Central to adaptive immunity is the interaction between the αβ T cell receptor (TCR) and peptide presented by the major histocompatibility complex (MHC) molecule. Presumably reflecting TCR-MHC bias and T cell signaling constraints, the TCR universally adopts a canonical polarity atop the MHC. We report the structures of two TCRs, derived from human induced T regulatory (iT(reg)) cells, complexed to an MHC class II molecule presenting a proinsulin-derived peptide. The ternary complexes revealed a 180° polarity reversal compared to all other TCR-peptide-MHC complex structures. Namely, the iT(reg) TCR α-chain and β-chain are overlaid with the α-chain and β-chain of MHC class II, respectively. Nevertheless, this TCR interaction elicited a peptide-reactive, MHC-restricted T cell signal. Thus TCRs are not 'hardwired' to interact with MHC molecules in a stereotypic manner to elicit a T cell signal, a finding that fundamentally challenges our understanding of TCR recognition.
- Infection and Immunity Program and The Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: