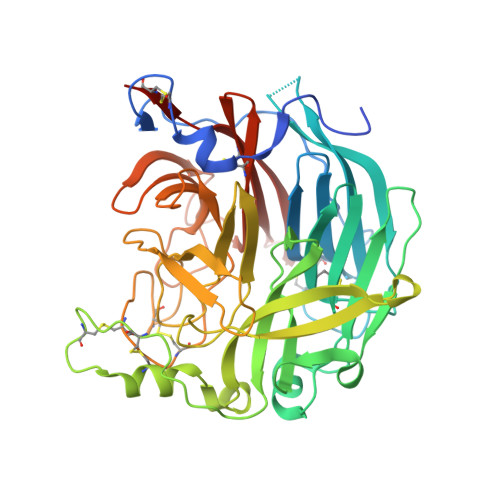
The catalytic mechanism of human parainfluenza virus type 3 haemagglutinin-neuraminidase revealed.
Dirr, L., El-Deeb, I.M., Guillon, P., Carroux, C.J., Chavas, L.M., von Itzstein, M.(2015) Angew Chem Int Ed Engl 54: 2936-2940
- PubMed: 25676091 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201412243
- Primary Citation Related Structures:
4XJQ, 4XJR - PubMed Abstract:
Human parainfluenza virus type 3 (hPIV-3) is one of the leading causes for lower respiratory tract disease in children, with neither an approved antiviral drug nor vaccine available to date. Understanding the catalytic mechanism of human parainfluenza virus haemagglutinin-neuraminidase (HN) protein is key to the design of specific inhibitors against this virus. Herein, we used (1) H NMR spectroscopy, X-ray crystallography, and virological assays to study the catalytic mechanism of the HN enzyme activity and have identified the conserved Tyr530 as a key amino acid involved in catalysis. A novel 2,3-difluorosialic acid derivative showed prolonged enzyme inhibition and was found to react and form a covalent bond with Tyr530. Furthermore, the novel derivative exhibited enhanced potency in virus blockade assays relative to its Neu2en analogue. These outcomes open the door for a new generation of potent inhibitors against hPIV-3 HN.
- Institute for Glycomics, Gold Coast Campus, Griffith University, Queensland, 4222 (Australia).
Organizational Affiliation: