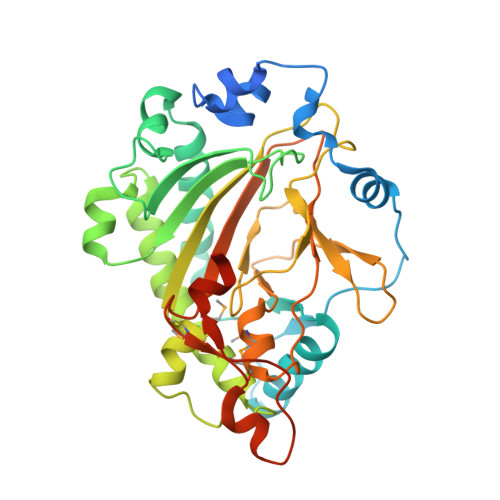
Structural Insights into Substrate Specificity of Feruloyl-CoA 6'-Hydroxylase from Arabidopsis thaliana.
Sun, X., Zhou, D., Kandavelu, P., Zhang, H., Yuan, Q., Wang, B.C., Rose, J., Yan, Y.(2015) Sci Rep 5: 10355-10355
- PubMed: 25993561 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep10355
- PubMed Abstract:
Coumarins belong to an important class of plant secondary metabolites. Feruloyl-CoA 6'-hydroxylase (F6'H), a 2-oxoglutarate dependent dioxygenase (2OGD), catalyzes a pivotal step in the biosynthesis of a simple coumarin scopoletin. In this study, we determined the 3-dimensional structure of the F6'H1 apo enzyme by X-ray crystallography. It is the first reported structure of a 2OGD enzyme involved in coumarin biosynthesis and closely resembles the structure of Arabidopsis thaliana anthocyanidin synthase. To better understand the mechanism of enzyme catalysis and substrate specificity, we also generated a homology model of a related ortho-hydroxylase (C2'H) from sweet potato. By comparing these two structures, we targeted two amino acid residues and verified their roles in substrate binding and specificity by site-directed mutagenesis.
- State Key Laboratory of Chemical Resource Engineering, Beijing University of Chemical Technology, Beijing 100029, China.
Organizational Affiliation: