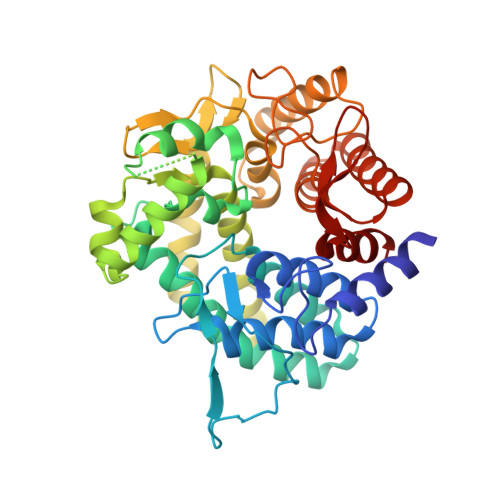
Crystal structure and substrate-binding mode of GH63 mannosylglycerate hydrolase from Thermus thermophilus HB8.
Miyazaki, T., Ichikawa, M., Iino, H., Nishikawa, A., Tonozuka, T.(2015) J Struct Biol 190: 21-30
- PubMed: 25712767 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2015.02.006
- Primary Citation Related Structures:
4WVA, 4WVB, 4WVC - PubMed Abstract:
Glycoside hydrolase family 63 (GH63) proteins are found in eukaryotes such as processing α-glucosidase I and also many bacteria and archaea. Recent studies have identified two bacterial and one plant GH63 mannosylglycerate hydrolases that act on both glucosylglycerate and mannosylglycerate, which are compatible solutes found in many thermophilic prokaryotes and some plants. Here we report the 1.67-Å crystal structure of one of these GH63 mannosylglycerate hydrolases, Tt8MGH from Thermus thermophilus HB8, which is 99% homologous to mannosylglycerate hydrolase from T. thermophilus HB27. Tt8MGH consists of a single (α/α)6-barrel catalytic domain with two additional helices and two long loops which form a homotrimer. The structures of this protein in complexes with glucose or glycerate were also determined at 1.77- or 2.10-Å resolution, respectively. A comparison of these structures revealed that the conformations of three flexible loops were largely different from each other. The conformational changes may be induced by ligand binding and serve to form finger-like structures for holding substrates. These findings represent the first-ever proposed substrate recognition mechanism for GH63 mannosylglycerate hydrolase.
- Department of Applied Biological Science, Tokyo University of Agriculture and Technology, 3-5-8 Saiwai-cho, Fuchu, Tokyo 183-8509, Japan.
Organizational Affiliation: