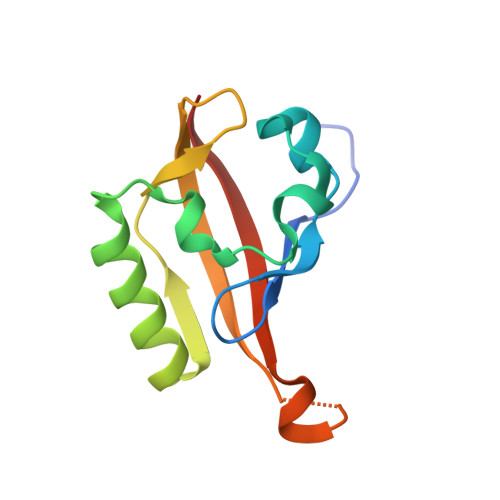
Unsaturated fatty acids as high-affinity ligands of the C-terminal Per-ARNT-Sim domain from the Hypoxia-inducible factor 3 alpha.
Fala, A.M., Oliveira, J.F., Adamoski, D., Aricetti, J.A., Dias, M.M., Dias, M.V.B., Sforca, M.L., Lopes-de-Oliveira, P.S., Rocco, S.A., Caldana, C., Dias, S.M.G., Ambrosio, A.L.B.(2015) Sci Rep 5: 12698-12698
- PubMed: 26237540 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep12698
- Primary Citation Related Structures:
4WN5 - PubMed Abstract:
Hypoxia-inducible transcription factors (HIF) form heterodimeric complexes that mediate cell responses to hypoxia. The oxygen-dependent stability and activity of the HIF-α subunits is traditionally associated to post-translational modifications such as hydroxylation, acetylation, ubiquitination, and phosphorylation. Here we report novel evidence showing that unsaturated fatty acids are naturally occurring, non-covalent structural ligands of HIF-3α, thus providing the initial framework for exploring its exceptional role as a lipid sensor under hypoxia.
- Laboratório Nacional de Biociências, Centro Nacional de Pesquisa em Energia e Materiais, Campinas, SP, Brazil, 13083-100.
Organizational Affiliation: