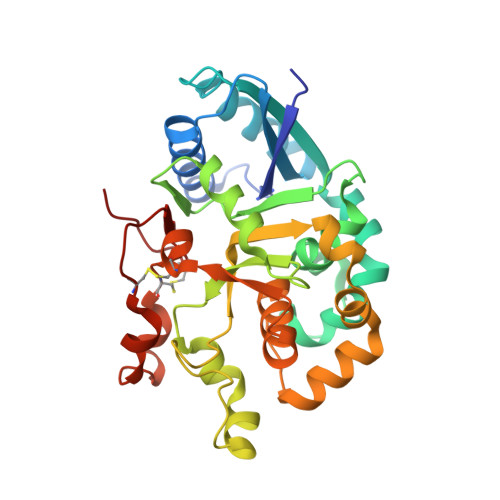
Notch-modifying xylosyltransferase structures support an SNi-like retaining mechanism.
Yu, H., Takeuchi, M., LeBarron, J., Kantharia, J., London, E., Bakker, H., Haltiwanger, R.S., Li, H., Takeuchi, H.(2015) Nat Chem Biol 11: 847-854
- PubMed: 26414444 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.1927
- Primary Citation Related Structures:
4WLG, 4WLM, 4WLZ, 4WM0, 4WMA, 4WMB, 4WMI, 4WMK, 4WN2 - PubMed Abstract:
A major question remaining in glycobiology is how a glycosyltransferase (GT) that retains the anomeric linkage of a sugar catalyzes the reaction. Xyloside α-1,3-xylosyltransferase (XXYLT1) is a retaining GT that regulates Notch receptor activation by adding xylose to the Notch extracellular domain. Here, using natural acceptor and donor substrates and active Mus musculus XXYLT1, we report a series of crystallographic snapshots along the reaction, including an unprecedented natural and competent Michaelis reaction complex for retaining enzymes. These structures strongly support the SNi-like reaction as the retaining mechanism for XXYLT1. Unexpectedly, the epidermal growth factor-like repeat acceptor substrate undergoes a large conformational change upon binding to the active site, providing a structural basis for substrate specificity. Our improved understanding of this retaining enzyme will accelerate the design of retaining GT inhibitors that can modulate Notch activity in pathological situations in which Notch dysregulation is known to cause cancer or developmental disorders.
- Biosciences Department, Brookhaven National Laboratory, Upton, New York, USA.
Organizational Affiliation: