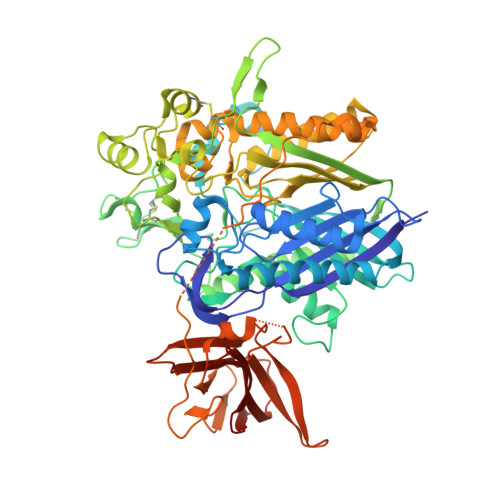
Structural Basis for Ceramide Recognition and Hydrolysis by Human Neutral Ceramidase.
Airola, M.V., Allen, W.J., Pulkoski-Gross, M.J., Obeid, L.M., Rizzo, R.C., Hannun, Y.A.(2015) Structure 23: 1482-1491
- PubMed: 26190575 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.06.013
- Primary Citation Related Structures:
4WGK - PubMed Abstract:
Neutral ceramidase (nCDase) catalyzes conversion of the apoptosis-associated lipid ceramide to sphingosine, the precursor for the proliferative factor sphingosine-1-phosphate. As an enzyme regulating the balance of ceramide and sphingosine-1-phosphate, nCDase is emerging as a therapeutic target for cancer. Here, we present the 2.6-Å crystal structure of human nCDase in complex with phosphate that reveals a striking, 20-Å deep, hydrophobic active site pocket stabilized by a eukaryotic-specific subdomain not present in bacterial ceramidases. Utilizing flexible ligand docking, we predict a likely binding mode for ceramide that superimposes closely with the crystallographically observed transition state analog phosphate. Our results suggest that nCDase uses a new catalytic strategy for Zn(2+)-dependent amidases, and generates ceramide specificity by sterically excluding sphingolipids with bulky headgroups and specifically recognizing the small hydroxyl head group of ceramide. Together, these data provide a foundation to aid drug development and establish common themes for how proteins recognize the bioactive lipid ceramide.
- Department of Medicine, Stony Brook University, Stony Brook, NY 11794, USA; Department of Medicine, Stony Brook Cancer Center, Stony Brook, NY 11794, USA.
Organizational Affiliation: