Picomolar Inhibitors of HIV-1 Reverse Transcriptase: Design and Crystallography of Naphthyl Phenyl Ethers.
Lee, W.G., Frey, K.M., Gallardo-Macias, R., Spasov, K.A., Bollini, M., Anderson, K.S., Jorgensen, W.L.(2014) ACS Med Chem Lett 5: 1259-1262
- PubMed: 25408842 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml5003713
- Primary Citation Related Structures:
4WE1 - PubMed Abstract:
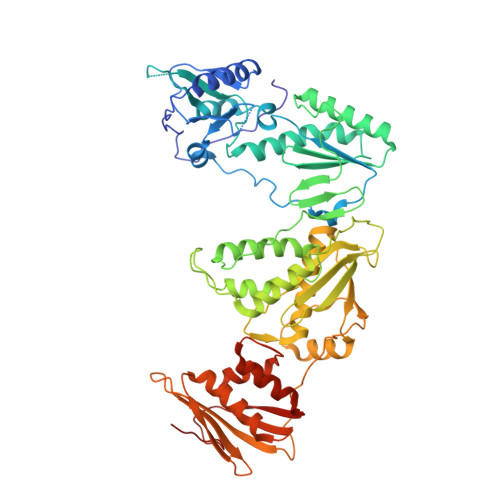
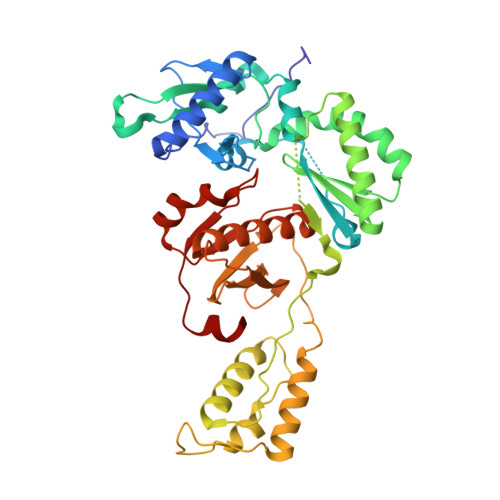
Catechol diethers that incorporate a 6-cyano-1-naphthyl substituent have been explored as non-nucleoside inhibitors of HIV-1 reverse transcriptase (NNRTIs). Promising compounds are reported that show midpicomolar activity against the wild-type virus and sub-20 nM activity against viral variants bearing Tyr181Cys and Lys103Asn mutations in HIV-RT. An X-ray crystal structure at 2.49 Å resolution is also reported for the key compound 6e with HIV-RT.
- Department of Chemistry, Yale University , New Haven, Connecticut 06520-8107, United States.
Organizational Affiliation: