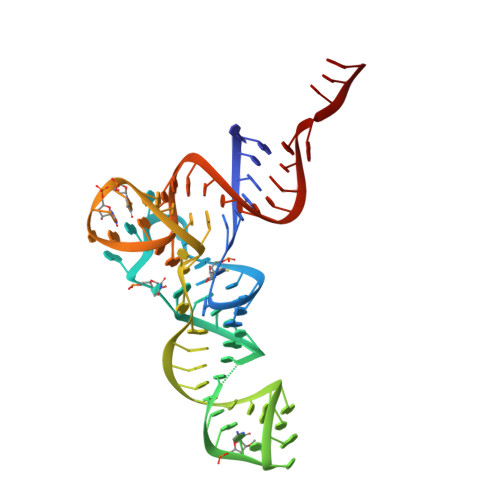
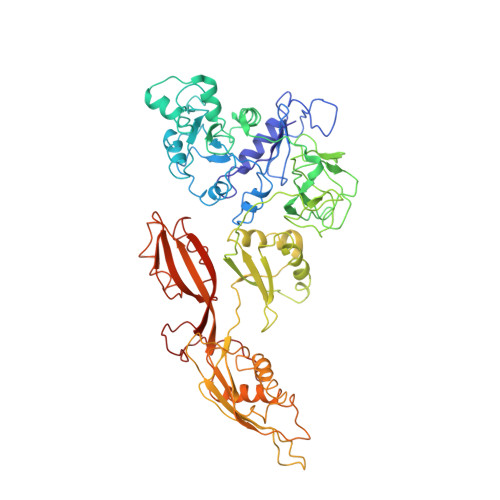
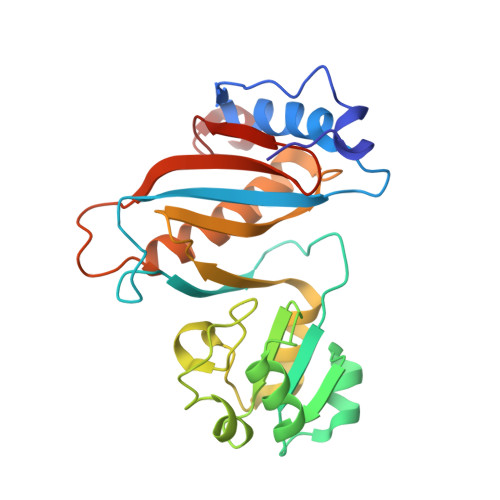
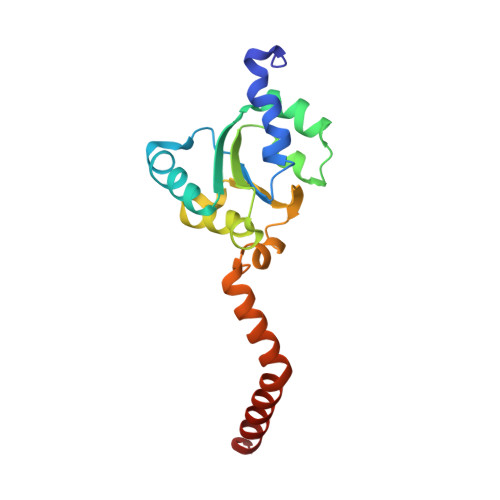
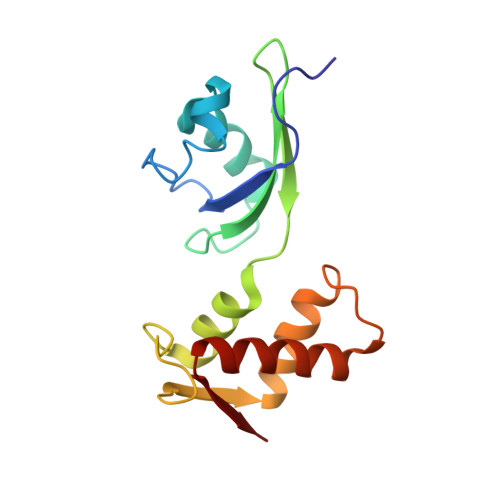
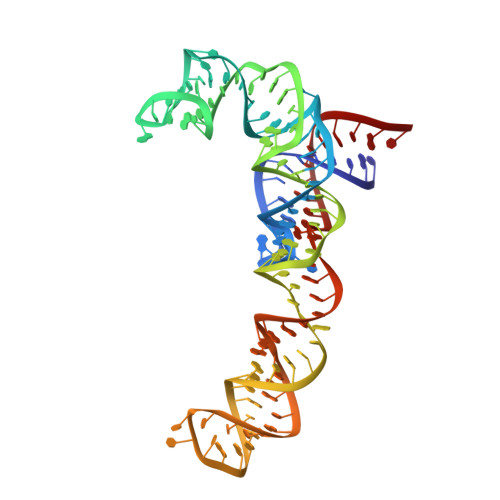
Mechanism of tetracycline resistance by ribosomal protection protein Tet(O).
Li, W., Atkinson, G.C., Thakor, N.S., Allas, U., Lu, C.C., Chan, K.Y., Tenson, T., Schulten, K., Wilson, K.S., Hauryliuk, V., Frank, J.(2013) Nat Commun 4: 1477-1477
- PubMed: 23403578 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms2470
- Primary Citation Related Structures:
4V6V - PubMed Abstract:
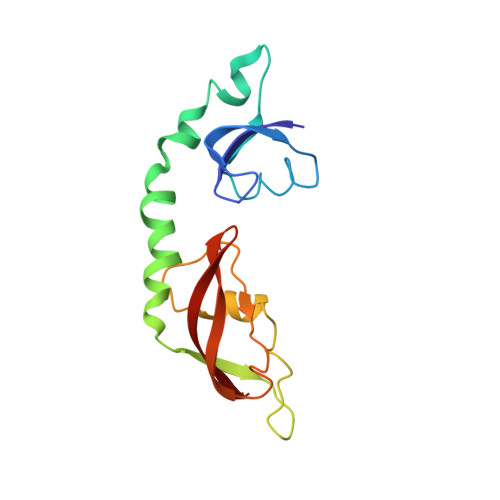
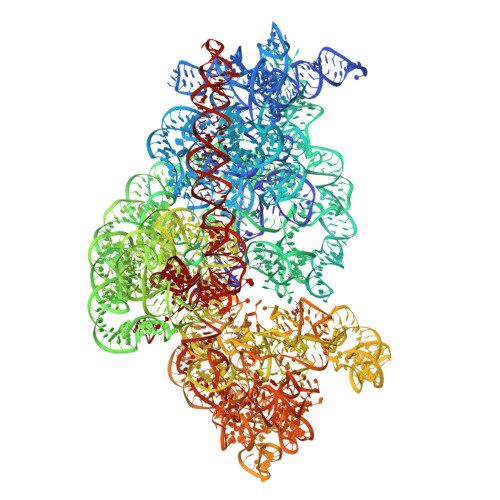
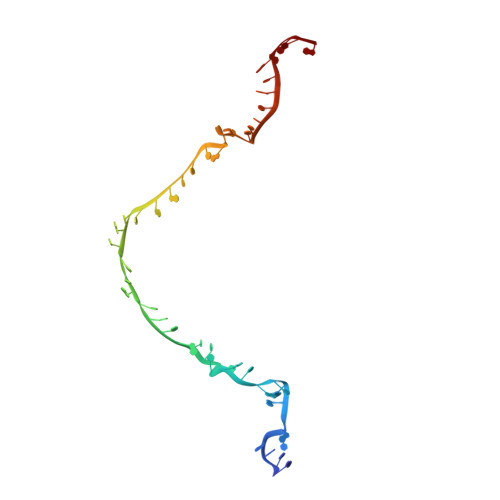
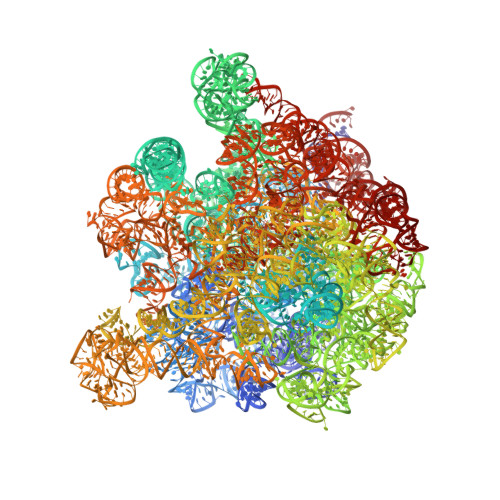
Tetracycline resistance protein Tet(O), which protects the bacterial ribosome from binding the antibiotic tetracycline, is a translational GTPase with significant similarity in both sequence and structure to the elongation factor EF-G. Here, we present an atomic model of the Tet(O)-bound 70S ribosome based on our cryo-electron microscopic reconstruction at 9.6-Å resolution. This atomic model allowed us to identify the Tet(O)-ribosome binding sites, which involve three characteristic loops in domain 4 of Tet(O). Replacements of the three amino-acid tips of these loops by a single glycine residue result in loss of Tet(O)-mediated tetracycline resistance. On the basis of these findings, the mechanism of Tet(O)-mediated tetracycline resistance can be explained in molecular detail.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, New York 10032, USA.
Organizational Affiliation: