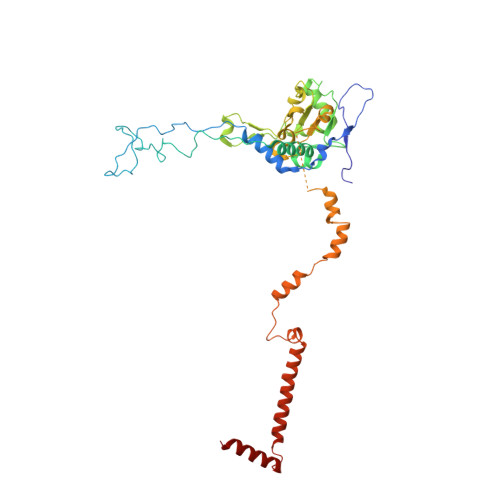
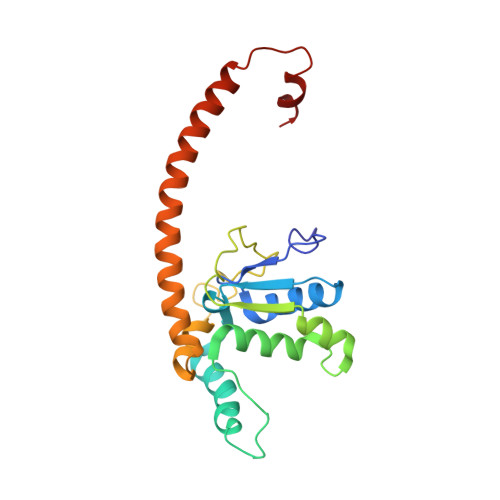
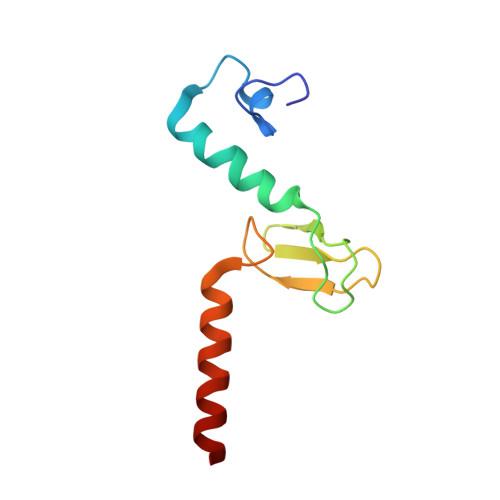
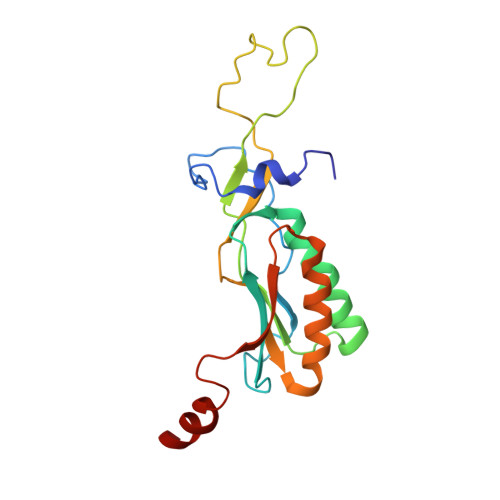
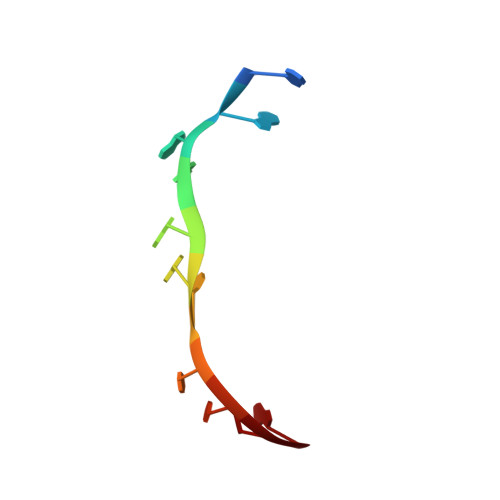
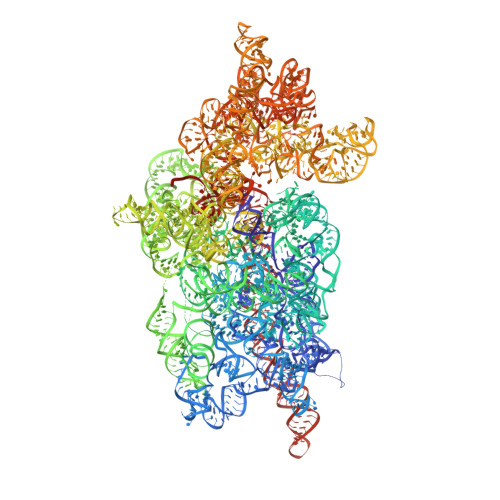
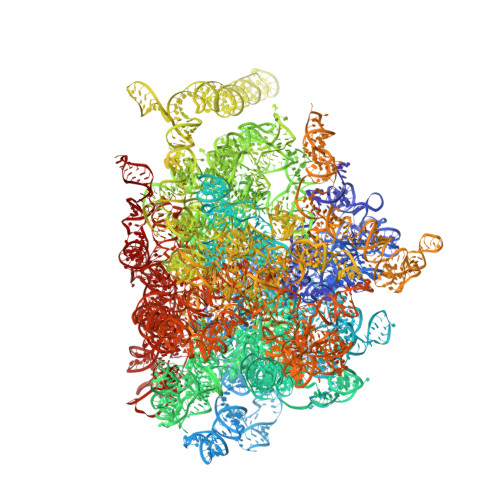
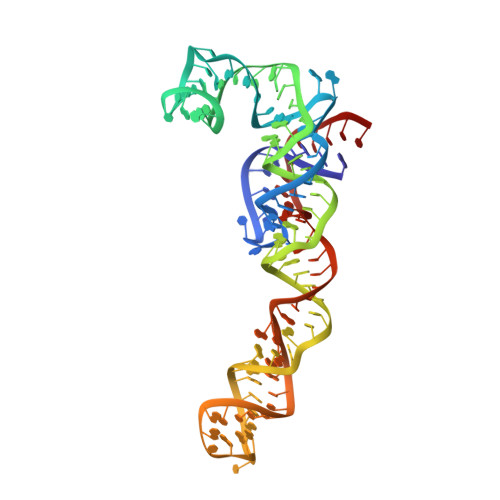
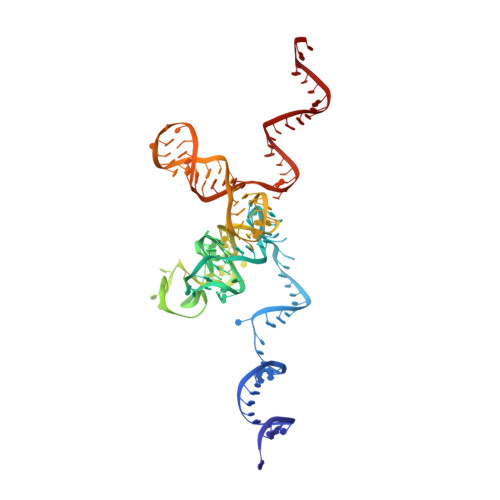
The molecular structure of the left-handed supra-molecular helix of eukaryotic polyribosomes.
Myasnikov, A.G., Afonina, Z.A., Menetret, J.F., Shirokov, V.A., Spirin, A.S., Klaholz, B.P.(2014) Nat Commun 5: 5294-5294
- PubMed: 25376914 Search on PubMed
- DOI: https://doi.org/10.1038/ncomms6294
- Primary Citation Related Structures:
4V3P - PubMed Abstract:
During protein synthesis, several ribosomes bind to a single messenger RNA (mRNA) forming large macromolecular assemblies called polyribosomes. Here we report the detailed molecular structure of a 100 MDa eukaryotic poly-ribosome complex derived from cryo electron tomography, sub-tomogram averaging and pseudo-atomic modelling by crystal structure fitting. The structure allowed the visualization of the three functional parts of the polysome assembly, the central core region that forms a rather compact left-handed supra-molecular helix, and the more open regions that harbour the initiation and termination sites at either ends. The helical region forms a continuous mRNA channel where the mRNA strand bridges neighbouring exit and entry sites of the ribosomes and prevents mRNA looping between ribosomes. This structure provides unprecedented insights into protein- and RNA-mediated inter-ribosome contacts that involve conserved sites through 40S subunits and long protruding RNA expansion segments, suggesting a role in stabilizing the overall polyribosomal assembly.
- 1] Centre for Integrative Biology (CBI), Department of Integrated Structural Biology, IGBMC (Institute of Genetics and of Molecular and Cellular Biology), 1 rue Laurent Fries, BP 10142, 67404 Illkirch, France [2] Centre National de la Recherche Scientifique (CNRS) UMR 7104, 67404 Illkirch, France [3] Institut National de la Santé et de la Recherche Médicale (INSERM), 67404 Illkirch, France [4] Université de Strasbourg, 67400 Strasbourg, France.
Organizational Affiliation: