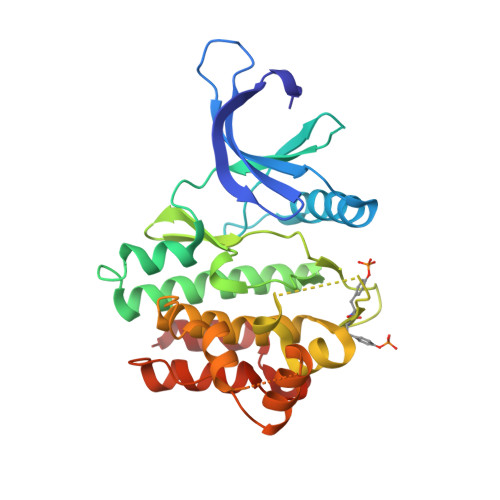
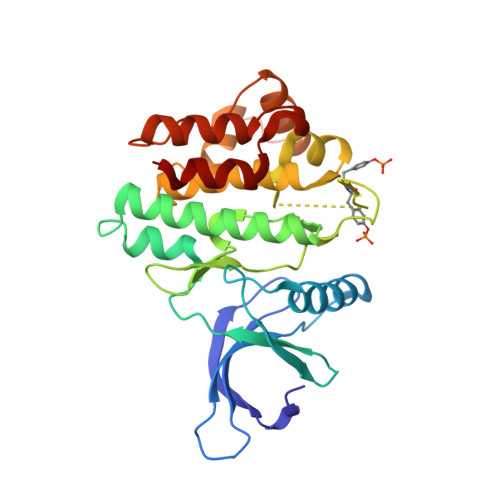
Utilisation of Structure Based Design to Identify Novel, Irreversible Inhibitors of the Epidermal Growth Factor Receptor (Egfr) Harboring the Gatekeeper T790M Mutation
Hennessy, E.J., Chuaquini, C., Ashton, S., Coclough, N., Cross, D.A.E., Debreczeni, J.E., Eberlein, C., Gingipalli, L., Klinowska, T.C.M., Orme, J.P., Sha, L., Wu, X.To be published.