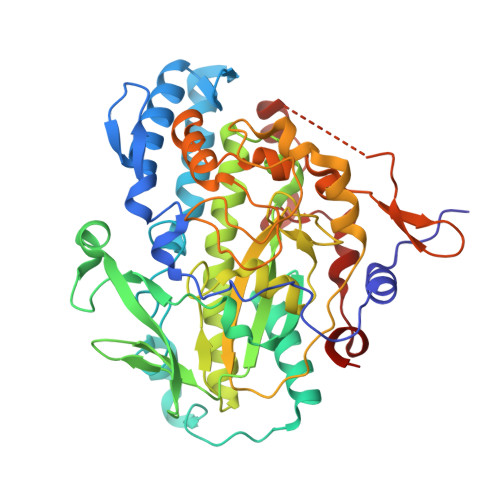
Structural Basis for Organohalide Respiration.
Bommer, M., Kunze, C., Fesseler, J., Schubert, T., Diekert, G., Dobbek, H.(2014) Science 346: 455
- PubMed: 25278505 Search on PubMed
- DOI: https://doi.org/10.1126/science.1258118
- Primary Citation Related Structures:
4UQU, 4UR0, 4UR1, 4UR2, 4UR3 - PubMed Abstract:
Organohalide-respiring microorganisms can use a variety of persistent pollutants, including trichloroethene (TCE), as terminal electron acceptors. The final two-electron transfer step in organohalide respiration is catalyzed by reductive dehalogenases. Here we report the x-ray crystal structure of PceA, an archetypal dehalogenase from Sulfurospirillum multivorans, as well as structures of PceA in complex with TCE and product analogs. The active site harbors a deeply buried norpseudo-B12 cofactor within a nitroreductase fold, also found in a mammalian B12 chaperone. The structures of PceA reveal how a cobalamin supports a reductive haloelimination exploiting a conserved B12-binding scaffold capped by a highly variable substrate-capturing region.
- Institut für Biologie, Strukturbiologie/Biochemie, Humboldt-Universität zu Berlin, Unter den Linden 6, 10099 Berlin, Germany.
Organizational Affiliation: