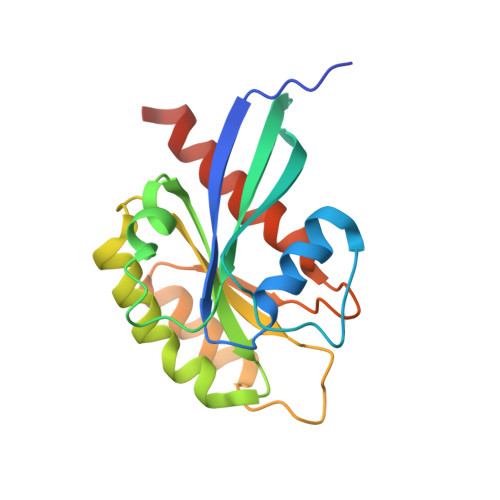
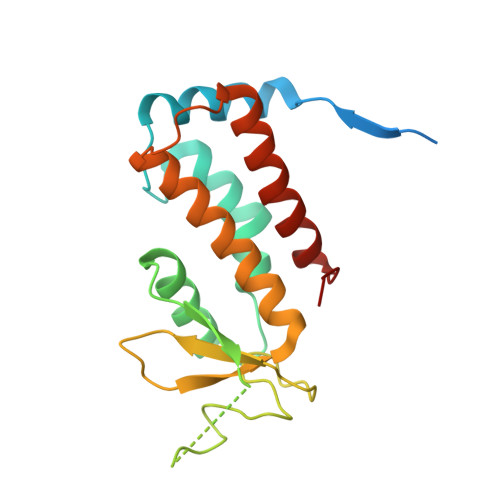
Structure of Rab11-Fip3-Rabin8 Reveals Simultaneous Binding of Fip3 and Rabin8 Effectors to Rab11.
Vetter, M., Stehle, R., Basquin, C., Lorentzen, E.(2015) Nat Struct Mol Biol 22: 695
- PubMed: 26258637 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3065
- Primary Citation Related Structures:
4UJ3, 4UJ4, 4UJ5 - PubMed Abstract:
The small GTPase Rab11 and its effectors FIP3 and Rabin8 are essential to membrane-trafficking pathways required for cytokinesis and ciliogenesis. Although effector binding is generally assumed to be sequential and mutually exclusive, we show that Rab11 can simultaneously bind FIP3 and Rabin8. We determined crystal structures of human Rab11-GMPPNP-Rabin8 and Rab11-GMPPNP-FIP3-Rabin8. The structures reveal that the C-terminal domain of Rabin8 adopts a previously undescribed fold that interacts with Rab11 at an unusual effector-binding site neighboring the canonical FIP3-binding site. We show that Rab11-GMPPNP-FIP3-Rabin8 is more stable than Rab11-GMPPNP-Rabin8, owing to direct interaction between Rabin8 and FIP3 within the dual effector-bound complex. The data allow us to propose a model for how membrane-targeting complexes assemble at the trans-Golgi network and recycling endosomes, through multiple weak interactions that create high-avidity complexes.
- Department of Structural Cell Biology, Max Planck Institute of Biochemistry, Martinsried, Germany.
Organizational Affiliation: