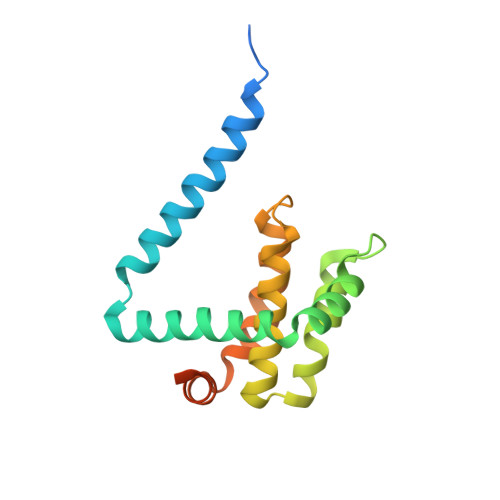
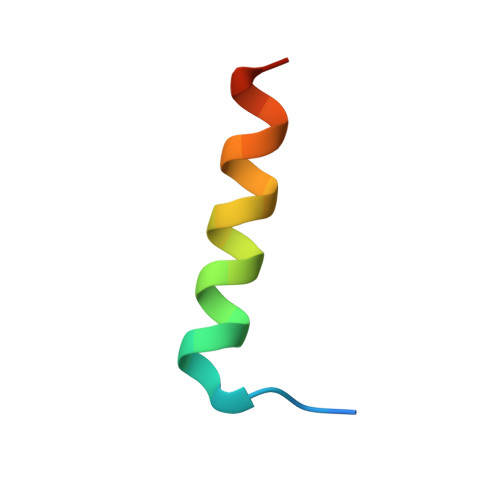
Structural Basis of Deerpox Virus-Mediated Inhibition of Apoptosis.
Burton, D.R., Caria, S., Marshall, B., Barry, M., Kvansakul, M.(2015) Acta Crystallogr D Biol Crystallogr 71: 1593
- PubMed: 26249341 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004715009402
- Primary Citation Related Structures:
4UF1, 4UF2, 4UF3 - PubMed Abstract:
Apoptosis is a key innate defence mechanism to eliminate virally infected cells. To counteract premature host-cell apoptosis, poxviruses have evolved numerous molecular strategies, including the use of Bcl-2 proteins, to ensure their own survival. Here, it is reported that the Deerpox virus inhibitor of apoptosis, DPV022, only engages a highly restricted set of death-inducing Bcl-2 proteins, including Bim, Bax and Bak, with modest affinities. Structural analysis reveals that DPV022 adopts a Bcl-2 fold with a dimeric domain-swapped topology and binds pro-death Bcl-2 proteins via two conserved ligand-binding grooves found on opposite sides of the dimer. Structures of DPV022 bound to Bim, Bak and Bax BH3 domains reveal that a partial obstruction of the binding groove is likely to be responsible for the modest affinities of DPV022 for BH3 domains. These findings reveal that domain-swapped dimeric Bcl-2 folds are not unusual and may be found more widely in viruses. Furthermore, the modest affinities of DPV022 for pro-death Bcl-2 proteins suggest that two distinct classes of anti-apoptotic viral Bcl-2 proteins exist: those that are monomeric and tightly bind a range of death-inducing Bcl-2 proteins, and others such as DPV022 that are dimeric and only bind a very limited number of death-inducing Bcl-2 proteins with modest affinities.
- Department of Biochemistry, La Trobe University, Melbourne, VIC 3058, Australia.
Organizational Affiliation: