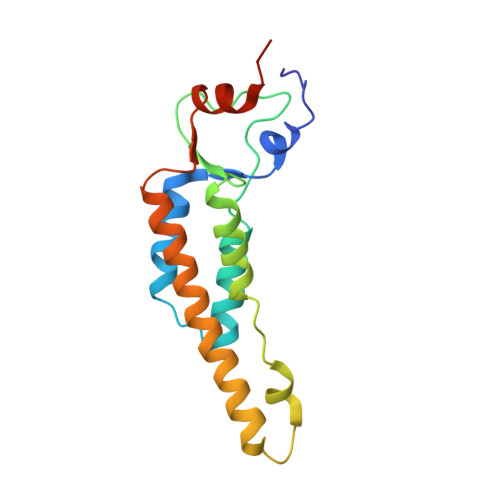
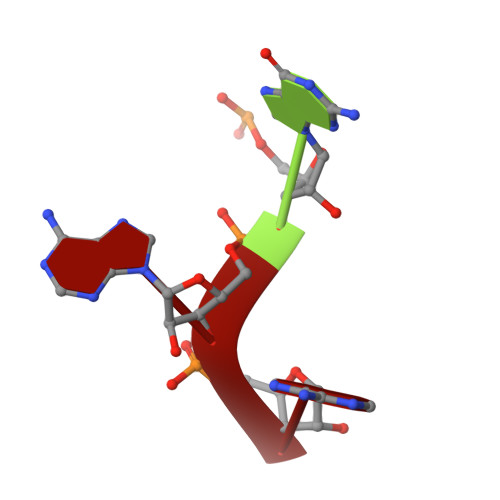
Seeing Tobacco Mosaic Virus Through Direct Electron Detectors.
Fromm, S.A., Bharat, T.A.M., Jakobi, A.J., Hagen, W.J.H., Sachse, C.(2015) J Struct Biol 189: 87
- PubMed: 25528571 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2014.12.002
- Primary Citation Related Structures:
4UDV - PubMed Abstract:
With the introduction of direct electron detectors (DED) to the field of electron cryo-microscopy, a wave of atomic-resolution structures has become available. As the new detectors still require comparative characterization, we have used tobacco mosaic virus (TMV) as a test specimen to study the quality of 3D image reconstructions from data recorded on the two direct electron detector cameras, K2 Summit and Falcon II. Using DED movie frames, we explored related image-processing aspects and compared the performance of micrograph-based and segment-based motion correction approaches. In addition, we investigated the effect of dose deposition on the atomic-resolution structure of TMV and show that radiation damage affects negative carboxyl chains first in a side-chain specific manner. Finally, using 450,000 asymmetric units and limiting the effects of radiation damage, we determined a high-resolution cryo-EM map at 3.35Å resolution. Here, we provide a comparative case study of highly ordered TMV recorded on different direct electron detectors to establish recording and processing conditions that enable structure determination up to 3.2Å in resolution using cryo-EM.
- EMBL - European Molecular Biology Laboratory, Structural and Computational Biology Unit, Meyerhofstr. 1, 69117 Heidelberg, Germany.
Organizational Affiliation: