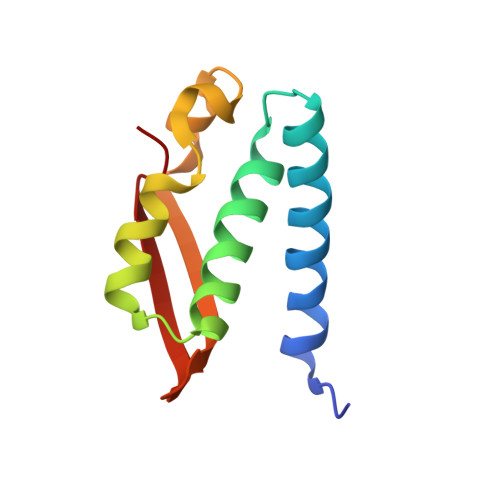
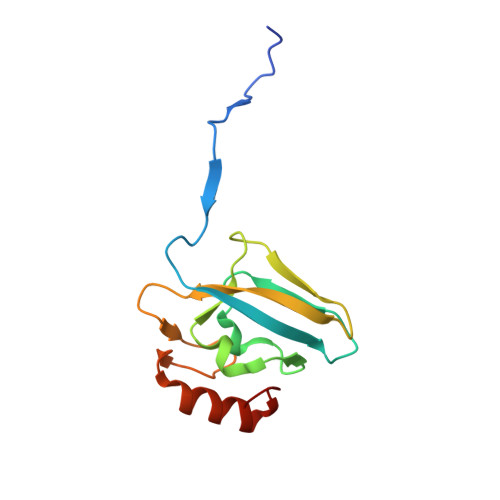
The complex of Bacillus pasteurii urease with acetohydroxamate anion from X-ray data at 1.55 A resolution.
Benini, S., Rypniewski, W.R., Wilson, K.S., Miletti, S., Ciurli, S., Mangani, S.(2000) J Biol Inorg Chem 5: 110-118
- PubMed: 10766443 Search on PubMed
- DOI: https://doi.org/10.1007/s007750050014
- Primary Citation Related Structures:
4UBP - PubMed Abstract:
The structure of Bacillus pasteurii urease inhibited with acetohydroxamic acid was solved and refined anisotropically using synchrotron X-ray cryogenic diffraction data (1.55 A resolution, 99.5% completeness, data redundancy = 26, R-factor = 15.1%, PDB code 4UBP). The two Ni ions in the active site are separated by a distance of 3.53 A. The structure clearly shows the binding mode of the inhibitor anion, symmetrically bridging the two Ni ions in the active site through the hydroxamate oxygen and chelating one Ni ion through the carbonyl oxygen. The flexible flap flanking the active site cavity is in the open conformation. The possible implications of the results on structure-based molecular design of new urease inhibitors are discussed.
- EMBL c/o DESY, Hamburg, Germany.
Organizational Affiliation: