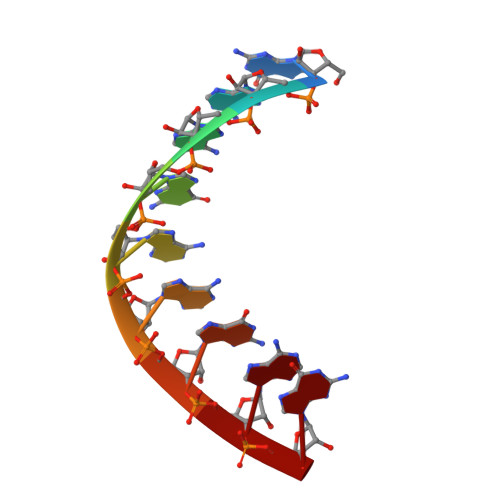
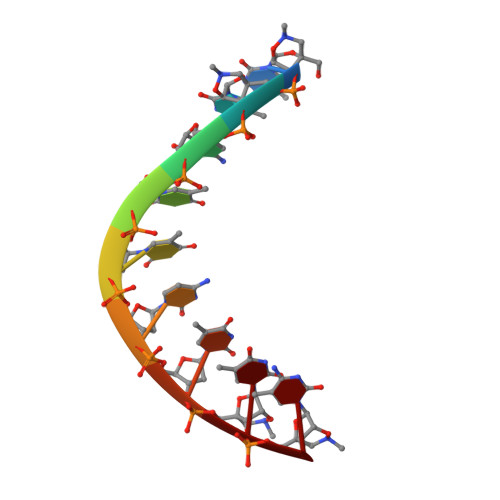
The crystal structure of a 2',4'-BNA(NC)[N-Me]-modified antisense gapmer in complex with the target RNA.
Kondo, J., Nomura, Y., Kitahara, Y., Obika, S., Torigoe, H.(2016) Chem Commun (Camb) 52: 2354-2357
- PubMed: 26731288 Search on PubMed
- DOI: https://doi.org/10.1039/c5cc08300a
- Primary Citation Related Structures:
4U6K, 4U6L, 4U6M - PubMed Abstract:
It has been confirmed by our previous studies that a 2',4'-BNA(NC)[N-Me]-modified antisense gapmer displays high affinity and selectivity to the target RNA strand, promising mRNA inhibitory activity and excellent nuclease resistance. Herein, we have obtained a crystal structure that provides insights into these excellent antisense properties.
- Department of Materials and Life Sciences, Sophia University, 7-1 Kioi-cho, Chiyoda-ku, 102-8554 Tokyo, Japan. j.kondo@sophia.ac.jp.
Organizational Affiliation: