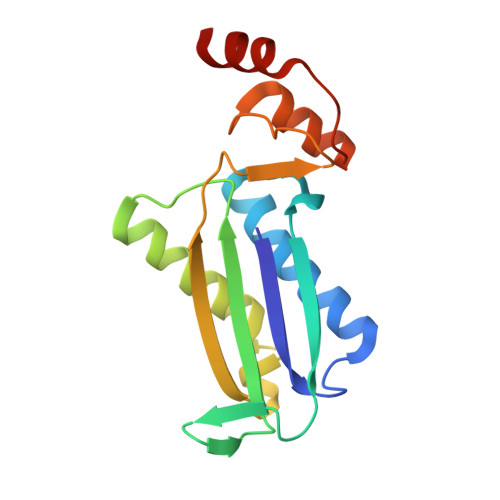
Functional and structural characterization of an unusual cofactor-independent oxygenase.
Baas, B.J., Poddar, H., Geertsema, E.M., Rozeboom, H.J., de Vries, M.P., Permentier, H.P., Thunnissen, A.M., Poelarends, G.J.(2015) Biochemistry 54: 1219-1232
- PubMed: 25565350 Search on PubMed
- DOI: https://doi.org/10.1021/bi501200j
- Primary Citation Related Structures:
4U5P, 4U5R - PubMed Abstract:
The vast majority of characterized oxygenases use bound cofactors to activate molecular oxygen to carry out oxidation chemistry. Here, we show that an enzyme of unknown activity, RhCC from Rhodococcus jostii RHA1, functions as an oxygenase, using 4-hydroxyphenylenolpyruvate as a substrate. This unique and complex reaction yields 3-hydroxy-3-(4-hydroxyphenyl)-pyruvate, 4-hydroxybenzaldehyde, and oxalic acid as major products. Incubations with H2(18)O, (18)O2, and a substrate analogue suggest that this enzymatic oxygenation reaction likely involves a peroxide anion intermediate. Analysis of sequence similarity and the crystal structure of RhCC (solved at 1.78 Å resolution) reveal that this enzyme belongs to the tautomerase superfamily. Members of this superfamily typically catalyze tautomerization, dehalogenation, or decarboxylation reactions rather than oxygenation reactions. The structure shows the absence of cofactors, establishing RhCC as a rare example of a redox-metal- and coenzyme-free oxygenase. This sets the stage to study the mechanistic details of cofactor-independent oxygen activation in the unusual context of the tautomerase superfamily.
- Department of Pharmaceutical Biology, Groningen Research Institute of Pharmacy and ‡Analytical Biochemistry and Mass Spectrometry Core Facility, Department of Pharmacy, University of Groningen , Antonius Deusinglaan 1, 9713 AV Groningen, The Netherlands.
Organizational Affiliation: