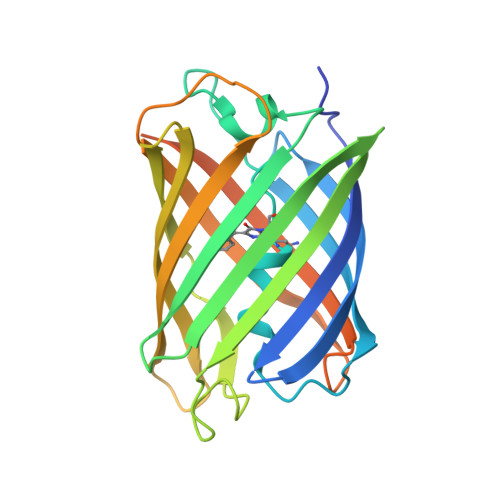
Thermal green protein, an extremely stable, nonaggregating fluorescent protein created by structure-guided surface engineering.
Close, D.W., Paul, C.D., Langan, P.S., Wilce, M.C., Traore, D.A., Halfmann, R., Rocha, R.C., Waldo, G.S., Payne, R.J., Rucker, J.B., Prescott, M., Bradbury, A.R.(2015) Proteins 83: 1225-1237
- PubMed: 25287913 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24699
- Primary Citation Related Structures:
4TZA, 4TZG - PubMed Abstract:
In this article, we describe the engineering and X-ray crystal structure of Thermal Green Protein (TGP), an extremely stable, highly soluble, non-aggregating green fluorescent protein. TGP is a soluble variant of the fluorescent protein eCGP123, which despite being highly stable, has proven to be aggregation-prone. The X-ray crystal structure of eCGP123, also determined within the context of this paper, was used to carry out rational surface engineering to improve its solubility, leading to TGP. The approach involved simultaneously eliminating crystal lattice contacts while increasing the overall negative charge of the protein. Despite intentional disruption of lattice contacts and introduction of high entropy glutamate side chains, TGP crystallized readily in a number of different conditions and the X-ray crystal structure of TGP was determined to 1.9 Å resolution. The structural reasons for the enhanced stability of TGP and eCGP123 are discussed. We demonstrate the utility of using TGP as a fusion partner in various assays and significantly, in amyloid assays in which the standard fluorescent protein, EGFP, is undesirable because of aberrant oligomerization.
- Bioscience Division, Los Alamos National Laboratory, Los Alamos, New Mexico.
Organizational Affiliation: