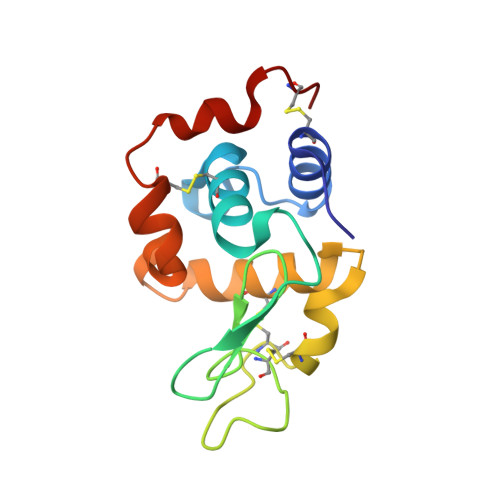
Biophysical aspects of lysozyme adduct with monocrotophos.
Amaraneni, S.R., Kumar, S., Gourinath, S.(2014) Anal Bioanal Chem 406: 5477-5485
- PubMed: 24969463 Search on PubMed
- DOI: https://doi.org/10.1007/s00216-014-7953-y
- Primary Citation Related Structures:
4TUN - PubMed Abstract:
The present study on in vitro formation and characterization of lysozyme adduct with monocrotophos (MP) evaluates the potential of lysozyme to be used as a sensitive biomarker to monitor exposure levels to the commonly used organophosphorus pesticide monocrotophos. Crystallization of lysozyme protein adduct with monocrotophos was also undertaken to understand the adduct formation mechanism at a molecular level. The binding of organophosphorus pesticides to lysozyme is one of the key steps in their mutagenicity. The formation and structural characterization of lysozyme adduct with monocrotophos was done using MALDI-TOFMS, fluorescence, UV/Vis spectroscopy, circular dichroism, and X-ray diffraction studies. We report the crystal structure of lysozyme adduct with monocrotophos at 1.9 Å. It crystallized in the P43 space group with two monomers in one asymmetric unit having one molecule of monocrotophos bound to each protein chain. The results proved that the fluorescence quenching of lysozyme by monocrotophos is due to binding of monocrotophos with a tryptophan residue of lysozyme. Monocrotophos interacts most strongly with the Trp-108 and Asp-52 of lysozyme. The interactions of the monocrotophos molecule with the lysozyme suggest the formation of a stable adduct. In addition, the alteration of lysozyme secondary structure in the presence of monocrotophos was confirmed by circular dichroism and fluorescence inhibition of lysozyme by increasing monocrotophos and UV/Vis spectrophotometry. The formation of lysozyme adduct with monocrotophos was confirmed by MALDI-TOFMS.
- Department of Chemistry, Alliance College of Engineering and Design, Alliance University, Bangalore, 562106, India, sramaraneni@yahoo.com.
Organizational Affiliation: