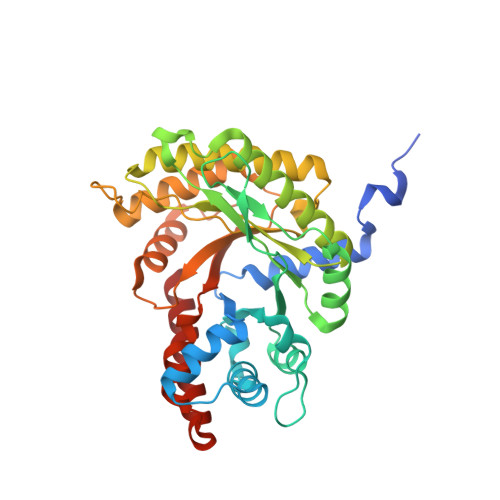
Structure of Toxoplasma gondii fructose-1,6-bisphosphate aldolase.
Boucher, L.E., Bosch, J.(2014) Acta Crystallogr F Struct Biol Commun 70: 1186-1192
- PubMed: 25195889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14017087
- Primary Citation Related Structures:
4TU1 - PubMed Abstract:
The apicomplexan parasite Toxoplasma gondii must invade host cells to continue its lifecycle. It invades different cell types using an actomyosin motor that is connected to extracellular adhesins via the bridging protein fructose-1,6-bisphosphate aldolase. During invasion, aldolase serves in the role of a structural bridging protein, as opposed to its normal enzymatic role in the glycolysis pathway. Crystal structures of the homologous Plasmodium falciparum fructose-1,6-bisphosphate aldolase have been described previously. Here, T. gondii fructose-1,6-bisphosphate aldolase has been crystallized in space group P22121, with the biologically relevant tetramer in the asymmetric unit, and the structure has been determined via molecular replacement to a resolution of 2.0 Å. An analysis of the quality of the model and of the differences between the four chains in the asymmetric unit and a comparison between the T. gondii and P. falciparum aldolase structures is presented.
- Department of Biochemistry and Molecular Biology, Johns Hopkins Bloomberg School of Public Health, 615 North Wolfe Street, Baltimore, MD 21205, USA.
Organizational Affiliation: