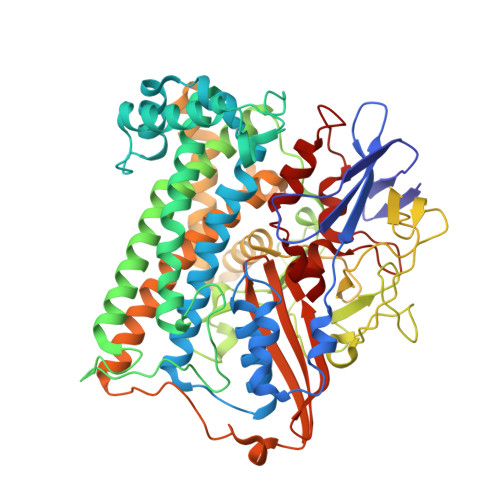
Tracking the route of molecular oxygen in O2-tolerant membrane-bound [NiFe] hydrogenase
Kalms, J., Schmidt, A., Frielingsdorf, S., Utesch, T., Gotthard, G., von Stetten, D., van der Linden, P., Royant, A., Mroginski, M., Carpentier, P., Lenz, O., Scheerer, P.(2018) Proc Natl Acad Sci U S A