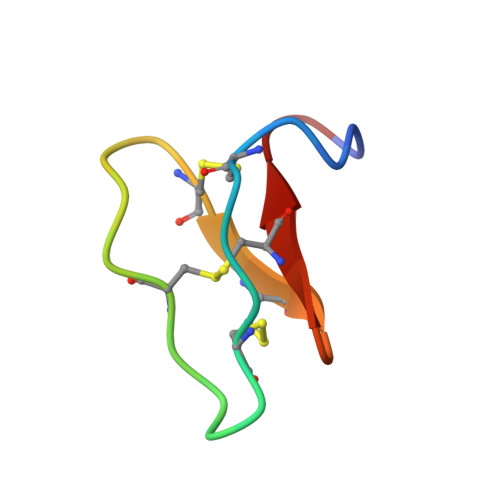
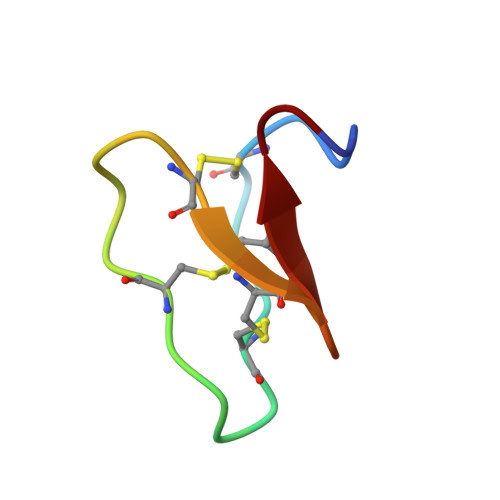
Racemic and Quasi-Racemic X-ray Structures of Cyclic Disulfide-Rich Peptide Drug Scaffolds.
Wang, C.K., King, G.J., Northfield, S.E., Ojeda, P.G., Craik, D.J.(2014) Angew Chem Int Ed Engl 53: 11236-11241
- PubMed: 25168664 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201406563
- Primary Citation Related Structures:
4TTK, 4TTL, 4TTM, 4TTN, 4TTO - PubMed Abstract:
Cyclic disulfide-rich peptides have exceptional stability and are promising frameworks for drug design. We were interested in obtaining X-ray structures of these peptides to assist in drug design applications, but disulfide-rich peptides can be notoriously difficult to crystallize. To overcome this limitation, we chemically synthesized the L- and D-forms of three prototypic cyclic disulfide-rich peptides: SFTI-1 (14-mer with one disulfide bond), cVc1.1 (22-mer with two disulfide bonds), and kB1 (29-mer with three disulfide bonds) for racemic crystallization studies. Facile crystal formation occurred from a racemic mixture of each peptide, giving structures solved at resolutions from 1.25 Å to 1.9 Å. Additionally, we obtained the quasi-racemic structures of two mutants of kB1, [G6A]kB1, and [V25A]kB1, which were solved at a resolution of 1.25 Å and 2.3 Å, respectively. The racemic crystallography approach appears to have broad utility in the structural biology of cyclic peptides.
- Institute for Molecular Bioscience, The University of Queensland, Brisbane, Qld, 4072 (Australia).
Organizational Affiliation: