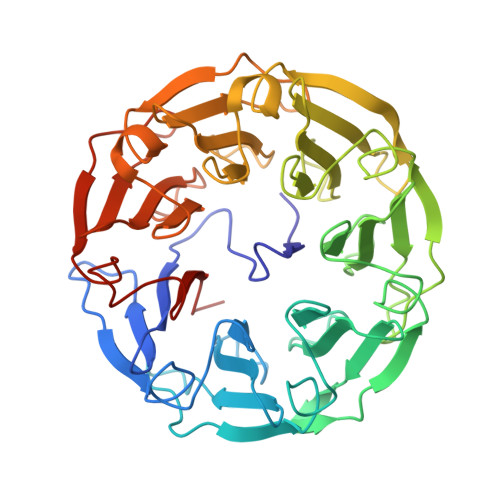
Structural Basis of Specific Recognition of Non-Reducing Terminal N-Acetylglucosamine by an Agrocybe aegerita Lectin.
Ren, X.M., Li, D.F., Jiang, S., Lan, X.Q., Hu, Y., Sun, H., Wang, D.C.(2015) PLoS One 10: e0129608-e0129608
- PubMed: 26114302 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0129608
- Primary Citation Related Structures:
4TQJ, 4TQK, 4TQM - PubMed Abstract:
O-linked N-acetylglucosaminylation (O-GlcNAcylation) is a reversible post-translational modification that plays essential roles in many cellular pathways. Research in this field, however, is hampered by the lack of suitable probes to identify, accumulate, and purify the O-GlcNAcylated proteins. We have previously reported the identification of a lectin from the mushroom Agrocybe aegerita, i.e., Agrocybe aegerita lectin 2, or AAL2, that could bind terminal N-acetylglucosamine with higher affinities and specificity than other currently used probes. In this paper, we report the crystal structures of AAL2 and its complexes with GlcNAc and GlcNAcβ1-3Galβ1-4GlcNAc and reveal the structural basis of GlcNAc recognition by AAL2 and residues essential for the binding of terminal N-acetylglucosamine. Study on AAL2 may enable us to design a protein probe that can be used to identify and purify O-GlcNAcylated proteins more efficiently.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Beijing, 100101, People's Republic of China; University of Chinese Academy of Sciences, No. 19A Yuquan Road, Beijing, 100049, People's Republic of China.
Organizational Affiliation: