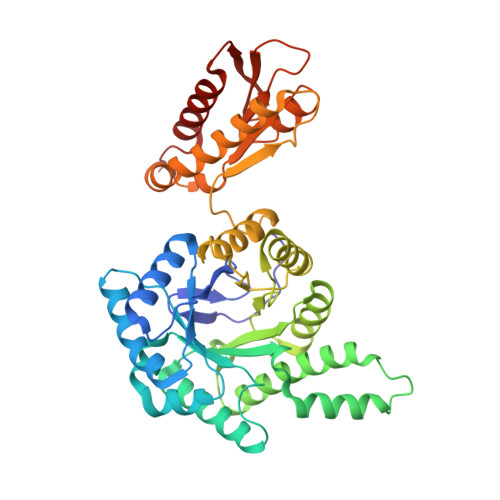
Structure of the GcpE-HMBPP complex from Thermus thermophilius.
Rekittke, I., Warkentin, E., Jomaa, H., Ermler, U.(2015) Biochem Biophys Res Commun 458: 246-250
- PubMed: 25660452 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2015.01.088
- Primary Citation Related Structures:
4S23 - PubMed Abstract:
Isoprenoid biosynthesis in many bacteria, plant chloroplasts and parasitic protozoa but not in humans proceeds via the mevalonate independent 2-C-methyl-D-erythritol-4-phosphate (MEP) pathway. Its penultimate reaction step is catalyzed by (E)-1-hydroxy-2-methyl-but-2-enyl-4-diphosphate (HMBPP) synthase (GcpE/IspG) which transforms 2-C-methyl-D-erythritol-2, 4-cyclo-diphosphate (MEcPP) to HMBPP. In this report we present the structure of GcpE of Thermus thermophiles in complex with its product HMBPP at a resolution of 1.65 Å. The GcpE-HMBPP like the GcpE-MEcPP structure is found in a closed, the ligand-free GcpE structure in an open enzyme state. Imposed by the rigid protein scaffold inside the active site funnel, linear HMBPP and circular MEcPP adopt highly similar conformations. The confined space also determines the conformational freedom of transition state intermediates and the design of anti-infective drugs. The apical Fe of the [4Fe-4S] cluster is coordinated to MEcPP in the GcpE-MEcPP complex and to a hydroxyl/water ligand but not to HMBPP in the GcpE-HMBPP complex. The GcpE-HMBPP structure can be attributed to one step in the currently proposed GcpE reaction cycle.
- Universitätsklinikum Gießen und Marburg GmbH, Marburg, Germany; Max-Planck-Institut für Biophysik, Frankfurt am Main, Germany.
Organizational Affiliation: