Crystal structure of Human Carnosinase-2 (CN2) in complex with inhibitor, Bestatin at 2.25 A
pandya, V., Kaushik, A., Singh, A.K., Singh, R.P., Kumaran, S.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cytosolic non-specific dipeptidase | 475 | Homo sapiens | Mutation(s): 0 Gene Names: CN2, CNDP dipeptidase 2 (metallopeptidase M20 family) [ Homo sapiens (human), CNDP2, CPGL, PEPA EC: 3.4.13.18 | 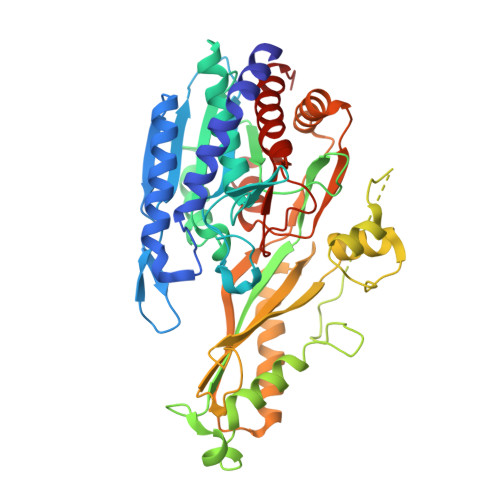 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q96KP4 GTEx: ENSG00000133313 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q96KP4 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BES Download:Ideal Coordinates CCD File | C [auth A], F [auth B] | 2-(3-AMINO-2-HYDROXY-4-PHENYL-BUTYRYLAMINO)-4-METHYL-PENTANOIC ACID C16 H24 N2 O4 VGGGPCQERPFHOB-RDBSUJKOSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | I [auth B], J [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| MN Download:Ideal Coordinates CCD File | D [auth A], E [auth A], G [auth B], H [auth B] | MANGANESE (II) ION Mn WAEMQWOKJMHJLA-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 87.09 | α = 90 |
| b = 100.07 | β = 90 |
| c = 105.83 | γ = 90 |
| Software Name | Purpose |
|---|---|
| MxCuBE | data collection |
| PHASES | phasing |
| PHENIX | refinement |
| iMOSFLM | data reduction |
| SCALA | data scaling |