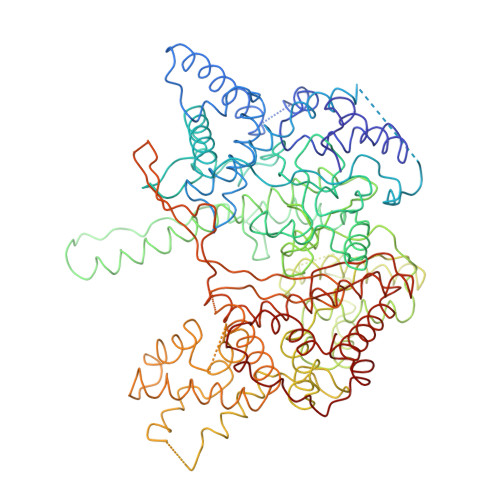
Crystal structure of bacteriophage T7 RNA polymerase at 3.3 A resolution.
Sousa, R., Chung, Y.J., Rose, J.P., Wang, B.C.(1993) Nature 364: 593-599
- PubMed: 7688864 Search on PubMed
- DOI: https://doi.org/10.1038/364593a0
- Primary Citation Related Structures:
4RNP - PubMed Abstract:
The crystal structure of T7 RNA polymerase reveals a molecule organized around a cleft that can accommodate a double-stranded DNA template. A portion (approximately 45%) of the molecule displays extensive structural homology to the polymerase domain of Klenow fragment and more limited homology to the human immunodeficiency virus HIV-1 reverse transcriptase. A comparison of the structures and sequences of these polymerases identifies structural elements that may be responsible for discriminating between ribonucleotide and deoxyribonucleotide substrates, and RNA and DNA templates. The relative locations of the catalytic site and a specific promoter recognition residue allow the orientation of the polymerase on the template to be defined.
- Department of Biological Sciences, University of Pittsburgh, Pennsylvania 15260.
Organizational Affiliation: