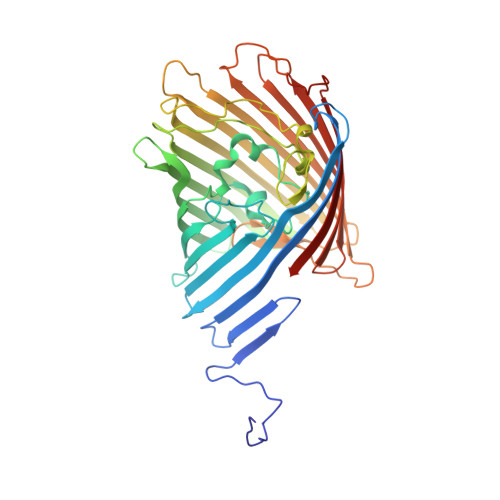
Structure, Dynamics, and Substrate Specificity of the OprO Porin from Pseudomonas aeruginosa.
Modi, N., Ganguly, S., Barcena-Uribarri, I., Benz, R., van den Berg, B., Kleinekathofer, U.(2015) Biophys J 109: 1429-1438
- PubMed: 26445443 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpj.2015.07.035
- Primary Citation Related Structures:
4RJW, 4RJX - PubMed Abstract:
The outer membrane (OM) of Gram-negative bacteria functions as a selective permeability barrier between cell and environment. For nutrient acquisition, the OM contains a number of channels that mediate uptake of small molecules by diffusion. Many of these channels are specific, i.e., they prefer certain substrates over others. In electrophysiological experiments, the OM channels OprP and OprO from Pseudomonas aeruginosa show a specificity for phosphate and diphosphate, respectively. In this study we use x-ray crystallography, free-energy molecular dynamics (MD) simulations, and electrophysiology to uncover the atomic basis for the different substrate specificity of these highly similar channels. A structural analysis of OprP and OprO revealed two crucial differences in the central constriction region. In OprP there are two tyrosine residues, Y62 and Y114, whereas the corresponding residues in OprO are phenylalanine F62 and aspartate D114. To probe the importance of these two residues in generating the different substrate specificities, the double mutants were generated in silico and in vitro. Applied-field MD simulations and electrophysiological experiments demonstrated that the double mutations interchange the phosphate and diphosphate specificities of OprP and OprO. Our findings outline a possible strategy to rationally design channel specificity by modification of a small number of residues that may be applicable to other pores as well.
- Department of Physics and Earth Sciences, Jacobs University Bremen, Bremen, Germany.
Organizational Affiliation: