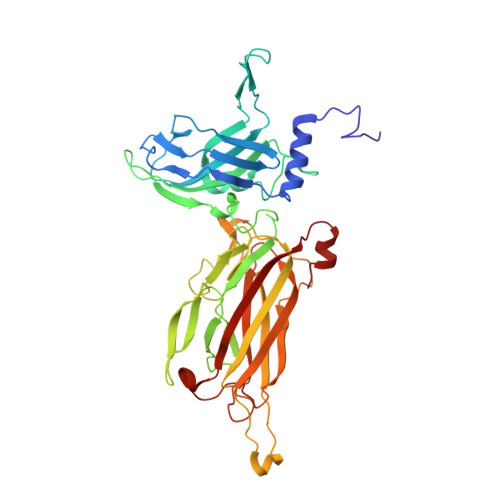
The targeted recognition of Lactococcus lactis phages to their polysaccharide receptors.
McCabe, O., Spinelli, S., Farenc, C., Labbe, M., Tremblay, D., Blangy, S., Oscarson, S., Moineau, S., Cambillau, C.(2015) Mol Microbiol 96: 875-886
- PubMed: 25708888 Search on PubMed
- DOI: https://doi.org/10.1111/mmi.12978
- Primary Citation Related Structures:
4RGA - PubMed Abstract:
Each phage infects a limited number of bacterial strains through highly specific interactions of the receptor-binding protein (RBP) at the tip of phage tail and the receptor at the bacterial surface. Lactococcus lactis is covered with a thin polysaccharide pellicle (hexasaccharide repeating units), which is used by a subgroup of phages as a receptor. Using L. lactis and phage 1358 as a model, we investigated the interaction between the phage RBP and the pellicle hexasaccharide of the host strain. A core trisaccharide (TriS), derived from the pellicle hexasaccharide repeating unit, was chemically synthesised, and the crystal structure of the RBP/TriS complex was determined. This provided unprecedented structural details of RBP/receptor site-specific binding. The complete hexasaccharide repeating unit was modelled and found to aptly fit the extended binding site. The specificity observed in in vivo phage adhesion assays could be interpreted in view of the reported structure. Therefore, by combining synthetic carbohydrate chemistry, X-ray crystallography and phage plaquing assays, we suggest that phage adsorption results from distinct recognition of the RBP towards the core TriS or the remaining residues of the hexasacchride receptor. This study provides a novel insight into the adsorption process of phages targeting saccharides as their receptors.
- Centre for Molecular Innovation and Drug Discovery, School of Chemistry and Chemical Biology, University College Dublin, Belfield, Dublin, Ireland.
Organizational Affiliation: