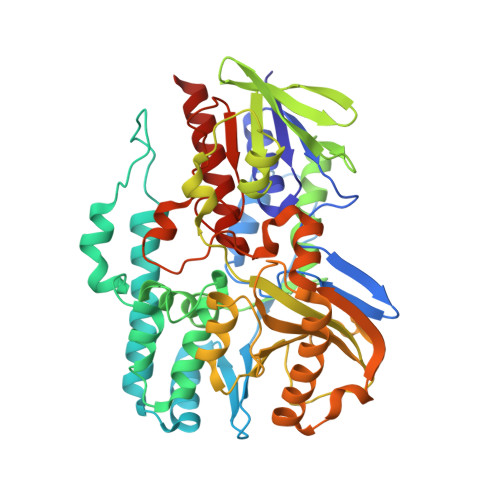
Crystal structure of 1'-OH-carotenoid 3,4-desaturase from Nonlabens dokdonensis DSW-6.
Ahn, J.W., Kim, K.J.(2015) Enzyme Microb Technol 77: 29-37
- PubMed: 26138397 Search on PubMed
- DOI: https://doi.org/10.1016/j.enzmictec.2015.05.005
- Primary Citation Related Structures:
4REP - PubMed Abstract:
The γ-carotenoids, such as myxol and saproxanthin, have a high potential to be utilized in nutraceutical and pharmaceutical industries for their neuro-protective and antioxidant effects. CrtD is involved in the production of γ-carotenoids by desaturating the C3'-C4' position of 1'-OH-γ-carotenoid. We determined the crystal structure of CrtD from Nonlabens dokdonensis DSW-6 (NdCrtD), the first structure of CrtD family enzymes. The NdCrtD structure was composed of two distinct domains, an FAD-binding domain and a substrate-binding domain, and the substrate-binding domain can be divided into two subdomains, a Rossmann fold-like subdomain and a lid subdomain. Although the FAD-binding domain showed a structure similar to canonical FAD-containing enzymes, the substrate-binding domain exhibited a novel structure to constitute a long and hydrophobic tunnel with a length of ∼40 Å. The molecular docking-simulation reveals that the tunnel provides an appropriate substrate-binding site for the carotenoid such as 1'-OH-γ-carotene with a length of ∼35 Å. We could predict residues related to recognize the 1'-hydroxyl group and to stabilize the hydrophobic end without hydroxyl group. Moreover, we suggest that the flexible entrance loop may undergo an open-closed formational change during the binding of the substrate.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu 702-701, South Korea.
Organizational Affiliation: