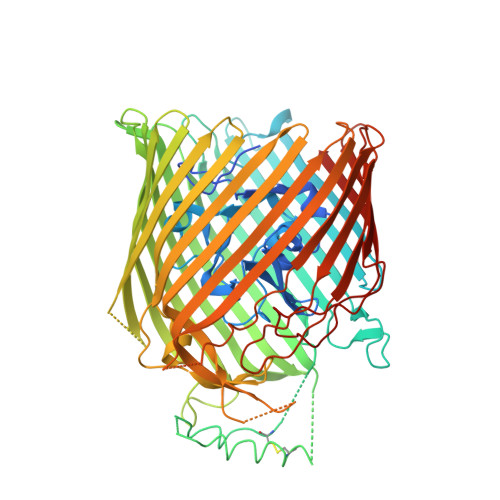
The molecular mechanism of Zinc acquisition by the neisserial outer-membrane transporter ZnuD.
Calmettes, C., Ing, C., Buckwalter, C.M., El Bakkouri, M., Chieh-Lin Lai, C., Pogoutse, A., Gray-Owen, S.D., Pomes, R., Moraes, T.F.(2015) Nat Commun 6: 7996-7996
- PubMed: 26282243 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms8996
- Primary Citation Related Structures:
4RDR, 4RDT, 4RVW - PubMed Abstract:
Invading bacteria from the Neisseriaceae, Acinetobacteriaceae, Bordetellaceae and Moraxellaceae families express the conserved outer-membrane zinc transporter zinc-uptake component D (ZnuD) to overcome nutritional restriction imposed by the host organism during infection. Here we demonstrate that ZnuD is required for efficient systemic infections by the causative agent of bacterial meningitis, Neisseria meningitidis, in a mouse model. We also combine X-ray crystallography and molecular dynamics simulations to gain insight into the mechanism of zinc recognition and transport across the bacterial outer-membrane by ZnuD. Because ZnuD is also considered a promising vaccine candidate against N. meningitidis, we use several ZnuD structural intermediates to map potential antigenic epitopes, and propose a mechanism by which ZnuD can maintain high sequence conservation yet avoid immune recognition by altering the conformation of surface-exposed loops.
- Department of Biochemistry, University of Toronto, 1 King's College Circle, Toronto, Ontario M5S 1A8, Canada.
Organizational Affiliation: