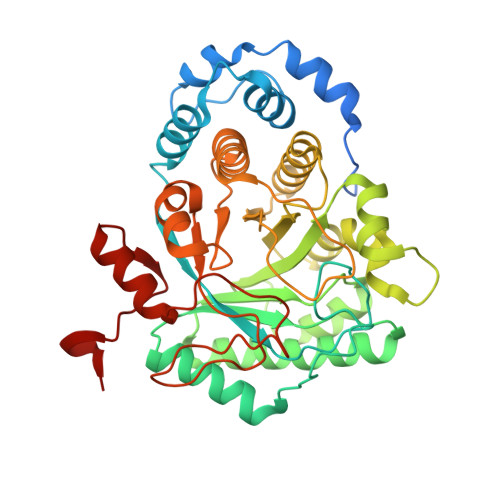
Crystal Structure of Tryptophan Lyase (NosL): Evidence for Radical Formation at the Amino Group of Tryptophan.
Nicolet, Y., Zeppieri, L., Amara, P., Fontecilla-Camps, J.C.(2014) Angew Chem Int Ed Engl 53: 11840-11844
- PubMed: 25196319 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201407320
- Primary Citation Related Structures:
4R33, 4R34 - PubMed Abstract:
Streptomyces actuosus tryptophan lyase (NosL) is a radical SAM enzyme which catalyzes the synthesis of 3-methyl-2-indolic acid, a precursor in the synthesis of the promising antibiotic nosiheptide. The reaction involves cleavage of the tryptophan Cα-Cβ bond and recombination of the amino-acid-derived -COOH fragment at the indole ring. Reported herein is the 1.8 Å resolution crystal structure of NosL complexed with its substrate. Unexpectedly, only one of the tryptophan amino hydrogen atoms is optimally placed for H abstraction by the SAM-derived 5'-deoxyadenosyl radical. This orientation, in turn, rules out the previously proposed delocalized indole radical as the species which undergoes Cα-Cβ bond cleavage. Instead, stereochemical considerations indicate that the reactive intermediate is a (·)NH tryptophanyl radical. A structure-based amino acid sequence comparison of NosL with the tyrosine lyases ThiH and HydG strongly suggests that an equivalent (·)NH radical operates in the latter enzymes.
- Metalloproteins Unit, Institut de Biologie Structurale, UMR5075, CEA, CNRS, Université Grenoble-Alpes, 71, avenue des Martyrs, CS 10090, 38044 Grenoble cedex 9 (France). yvain.nicolet@ibs.fr.
Organizational Affiliation: