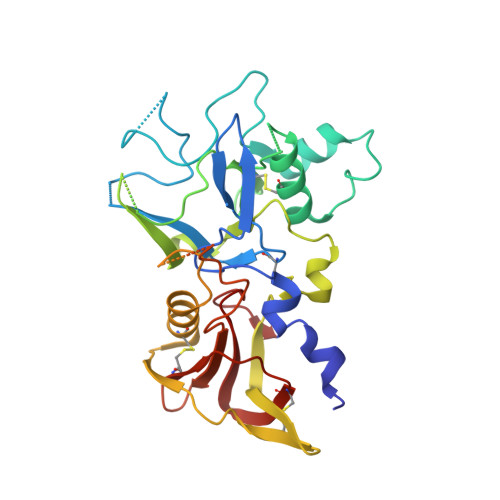
Structure and Dynamics of Apical Membrane Antigen 1 from Plasmodium falciparum FVO.
Lim, S.S., Yang, W., Krishnarjuna, B., Kannan Sivaraman, K., Chandrashekaran, I.R., Kass, I., MacRaild, C.A., Devine, S.M., Debono, C.O., Anders, R.F., Scanlon, M.J., Scammells, P.J., Norton, R.S., McGowan, S.(2014) Biochemistry 53: 7310-7320
- PubMed: 25360546 Search on PubMed
- DOI: https://doi.org/10.1021/bi5012089
- Primary Citation Related Structures:
4R19, 4R1A, 4R1B, 4R1C - PubMed Abstract:
Apical membrane antigen 1 (AMA1) interacts with RON2 to form a protein complex that plays a key role in the invasion of host cells by malaria parasites. Blocking this protein-protein interaction represents a potential route to controlling malaria and related parasitic diseases, but the polymorphic nature of AMA1 has proven to be a major challenge to vaccine-induced antibodies and peptide inhibitors exerting strain-transcending inhibitory effects. Here we present the X-ray crystal structure of AMA1 domains I and II from Plasmodium falciparum strain FVO. We compare our new structure to those of AMA1 from P. falciparum 3D7 and Plasmodium vivax. A combination of normalized B factor analysis and computational methods has been used to investigate the flexibility of the domain I loops and how this correlates with their roles in determining the strain specificity of human antibody responses and inhibitory peptides. We also investigated the domain II loop, a key region involved in inhibitor binding, by comparison of multiple AMA1 crystal structures. Collectively, these results provide valuable insights that should contribute to the design of strain-transcending agents targeting P. falciparum AMA1.
- Medicinal Chemistry, Monash Institute of Pharmaceutical Sciences, Monash University , Parkville, Victoria 3052, Australia.
Organizational Affiliation: