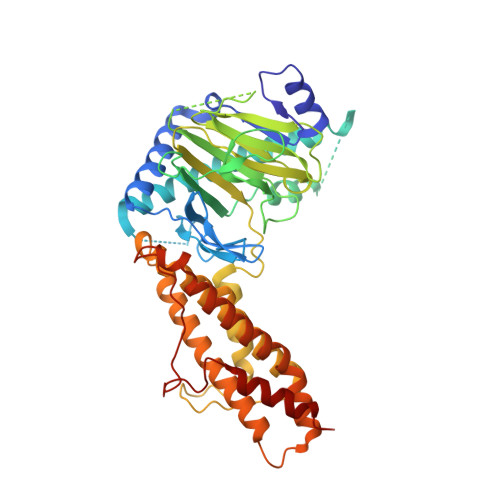
Meclofenamic acid selectively inhibits FTO demethylation of m6A over ALKBH5.
Huang, Y., Yan, J., Li, Q., Li, J., Gong, S., Zhou, H., Gan, J., Jiang, H., Jia, G.F., Luo, C., Yang, C.G.(2015) Nucleic Acids Res 43: 373-384
- PubMed: 25452335 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku1276
- Primary Citation Related Structures:
4QKN - PubMed Abstract:
Two human demethylases, the fat mass and obesity-associated (FTO) enzyme and ALKBH5, oxidatively demethylate abundant N(6)-methyladenosine (m(6)A) residues in mRNA. Achieving a method for selective inhibition of FTO over ALKBH5 remains a challenge, however. Here, we have identified meclofenamic acid (MA) as a highly selective inhibitor of FTO. MA is a non-steroidal, anti-inflammatory drug that mechanistic studies indicate competes with FTO binding for the m(6)A-containing nucleic acid. The structure of FTO/MA has revealed much about the inhibitory function of FTO. Our newfound understanding, revealed herein, of the part of the nucleotide recognition lid (NRL) in FTO, for example, has helped elucidate the principles behind the selectivity of FTO over ALKBH5. Treatment of HeLa cells with the ethyl ester form of MA (MA2) has led to elevated levels of m(6)A modification in mRNA. Our collective results highlight the development of functional probes of the FTO enzyme that will (i) enable future biological studies and (ii) pave the way for the rational design of potent and specific inhibitors of FTO for use in medicine.
- CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China.
Organizational Affiliation: