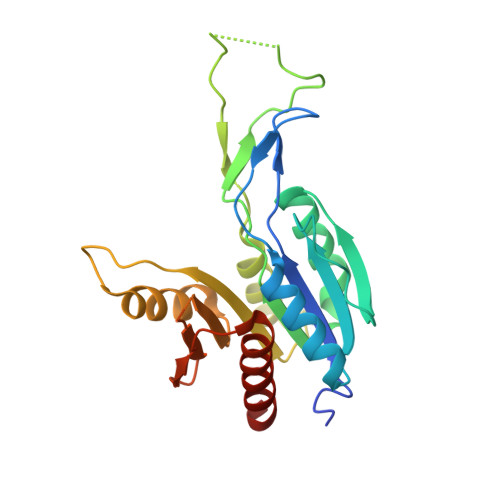
Crystal Structure of the C-terminal CTP-binding domain of a Phosphopantothenoylcysteine decarboxylase/phosphopantothenate-cysteine ligase with bound CTP from Mycobacterium smegmatis
Dranow, D.M., Abendroth, J., Wernimont, A.K., Edwards, T.E., Lorimer, D.To be published.