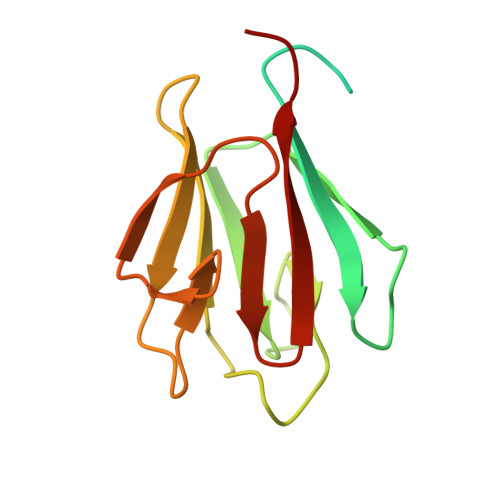
Interaction of 2-oxoglutarate dehydrogenase OdhA with its inhibitor OdhI in Corynebacterium glutamicum: Mutants and a model.
Raasch, K., Bocola, M., Labahn, J., Leitner, A., Eggeling, L., Bott, M.(2014) J Biotechnol 191: 99-105
- PubMed: 24905147 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbiotec.2014.05.023
- Primary Citation Related Structures:
4QCJ - PubMed Abstract:
Pyruvate dehydrogenase and oxoglutarate dehydrogenase catalyze key reactions in central metabolism. In Corynebacterium glutamicum and related bacteria like Mycobacterium tuberculosis both activities reside in a novel protein supercomplex with the fusion protein OdhA catalyzing the conversion of oxoglutarate to succinyl-coenzyme A. This activity is inhibited by the forkhead-associated (FHA) domain of the small autoinhibitory protein OdhI. Here we used a biological screen which enabled us to isolate suppressor mutants that are influenced in OdhA-OdhI interaction. Five mutants carrying an OdhI mutation were isolated and one with an OdhA mutation. The OdhA mutein OdhA-C704E and three additional C704 variants were constructed. They exhibited unaltered or even slightly enhanced OdhA activity but showed reduced inhibition and interaction with OdhI. The FHA domain of OdhI was crystallized and its structure found in full agreement with previously determined NMR structures. Based on further structural studies, OdhA-OdhI crosslinking experiments, and modeling we discuss the experimental data generated on OdhA-OdhI interaction, with the latter protein representing a rare example of an FHA domain also recognizing a non-phosphorylated interaction partner.
- Institute of Bio- and Geosciences, IBG-1: Biotechnology, Forschungszentrum Jülich, Jülich, Germany.
Organizational Affiliation: