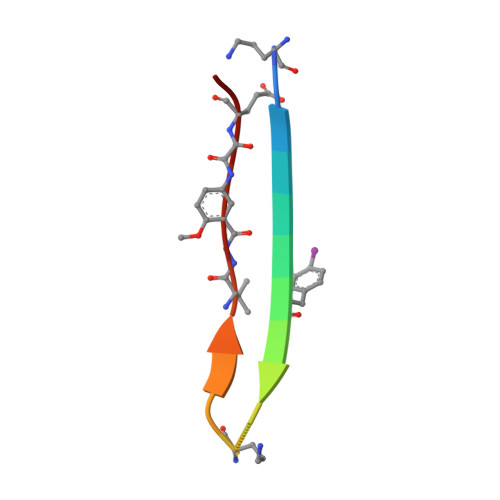
A Fibril-Like Assembly of Oligomers of a Peptide Derived from beta-Amyloid.
Pham, J.D., Spencer, R.K., Chen, K.H., Nowick, J.S.(2014) J Am Chem Soc 136: 12682-12690
- PubMed: 25068693 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja505713y
- Primary Citation Related Structures:
4Q8D - PubMed Abstract:
A macrocyclic β-sheet peptide containing two nonapeptide segments based on Aβ(15-23) (QKLVFFAED) forms fibril-like assemblies of oligomers in the solid state. The X-ray crystallographic structure of macrocyclic β-sheet peptide 3 was determined at 1.75 Å resolution. The macrocycle forms hydrogen-bonded dimers, which further assemble along the fibril axis in a fashion resembling a herringbone pattern. The extended β-sheet comprising the dimers is laminated against a second layer of dimers through hydrophobic interactions to form a fibril-like assembly that runs the length of the crystal lattice. The second layer is offset by one monomer subunit, so that the fibril-like assembly is composed of partially overlapping dimers, rather than discrete tetramers. In aqueous solution, macrocyclic β-sheet 3 and homologues 4 and 5 form discrete tetramers, rather than extended fibril-like assemblies. The fibril-like assemblies of oligomers formed in the solid state by macrocyclic β-sheet 3 represent a new mode of supramolecular assembly not previously observed for the amyloidogenic central region of Aβ. The structures observed at atomic resolution for this peptide model system may offer insights into the structures of oligomers and oligomer assemblies formed by full-length Aβ and may provide a window into the propagation and replication of amyloid oligomers.
- Department of Chemistry, University of California, Irvine , Irvine, California 92697-2025, United States.
Organizational Affiliation: