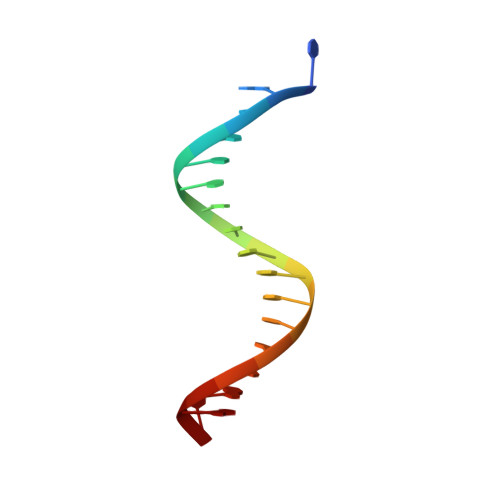
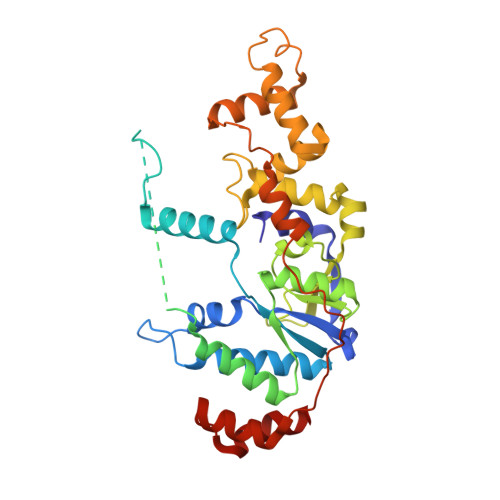
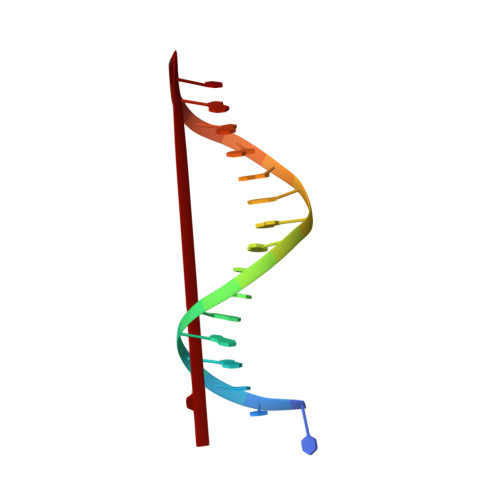
Crystal structure of the catalytic core of Rad2: insights into the mechanism of substrate binding.
Mietus, M., Nowak, E., Jaciuk, M., Kustosz, P., Studnicka, J., Nowotny, M.(2014) Nucleic Acids Res 42: 10762-10775
- PubMed: 25120270 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku729
- Primary Citation Related Structures:
4Q0R, 4Q0W, 4Q0Z, 4Q10 - PubMed Abstract:
Rad2/XPG belongs to the flap nuclease family and is responsible for a key step of the eukaryotic nucleotide excision DNA repair (NER) pathway. To elucidate the mechanism of DNA binding by Rad2/XPG, we solved crystal structures of the catalytic core of Rad2 in complex with a substrate. Rad2 utilizes three structural modules for recognition of the double-stranded portion of DNA substrate, particularly a Rad2-specific α-helix for binding the cleaved strand. The protein does not specifically recognize the single-stranded portion of the nucleic acid. Our data suggest that in contrast to related enzymes (FEN1 and EXO1), the Rad2 active site may be more accessible, which would create an exit route for substrates without a free 5' end.
- Laboratory of Protein Structure, International Institute of Molecular and Cell Biology, Warsaw 02-109, Poland.
Organizational Affiliation: